Journal Description
Applied Microbiology
Applied Microbiology
is an international, peer-reviewed, open access journal on application of microorganisms published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 13.3 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- Applied Microbiology is a companion journal of Microorganisms.
Latest Articles
Effect of kuratsuki Bacillus and Priestia on Taste of Sake
Appl. Microbiol. 2024, 4(1), 147-161; https://doi.org/10.3390/applmicrobiol4010011 - 15 Jan 2024
Abstract
The co-cultivation of sake yeast (AK25, K901, K1401, or K1801 strain) and the kuratsuki Bacillus A-10 and/or Priestia B-12 strains in koji solution was performed to demonstrate the effects of these two kuratsuki bacteria on sake taste. The results showed that the Brix
[...] Read more.
The co-cultivation of sake yeast (AK25, K901, K1401, or K1801 strain) and the kuratsuki Bacillus A-10 and/or Priestia B-12 strains in koji solution was performed to demonstrate the effects of these two kuratsuki bacteria on sake taste. The results showed that the Brix and acidity patterns of sake preparations produced with and without these kuratsuki bacteria were very similar. This indicated that the addition of these kuratsuki bacteria did not inhibit ethanol fermentation or organic acid production by sake yeast. A taste recognition device showed that the effects of these kuratsuki bacteria on the saltiness and sourness of sake were greater than those on other taste properties. Astringency stimulation and saltiness of sake produced using the sake yeast K901 were increased by Bacillus A-10 and decreased by Priestia B-12. Except for these two cases, the taste intensities of sake preparations produced with the Bacillus A-10 and Priestia B-12 strains were very similar, but differed from those of sake produced with kuratsuki Kocuria. These results support our hypothesis that the flavor and taste of sake can be controlled by utilizing the interactions between kuratsuki bacteria and sake yeast. For crating the desired sake taste, a combination of kuratsuki bacteria and sake yeast should be considered.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
Open AccessArticle
Genetic Analysis and Epidemiological Impact of SARS-CoV-2: A Multinational Study of 1000 Samples Using RT-PCR
by
, , , , , , , and
Appl. Microbiol. 2024, 4(1), 133-146; https://doi.org/10.3390/applmicrobiol4010010 - 15 Jan 2024
Abstract
►▼
Show Figures
The ongoing global public health challenge posed by the COVID-19 pandemic necessitates continuous research and surveillance efforts. In this study, we comprehensively analyzed over 1000 COVID-19 RT-PCR tests conducted on a cohort of 1200 patients in Saudi Arabia. Our primary goal was to
[...] Read more.
The ongoing global public health challenge posed by the COVID-19 pandemic necessitates continuous research and surveillance efforts. In this study, we comprehensively analyzed over 1000 COVID-19 RT-PCR tests conducted on a cohort of 1200 patients in Saudi Arabia. Our primary goal was to investigate mutations in specific genes RdRp, N, and E different infection and recovery stages in Saudi patients with SARS-CoV-2. We also extended our analysis to include patients of various nationalities residing in Saudi Arabia, with the overarching objective of assessing these genes as markers for COVID-19 presence and progression. To diagnose and investigate potential genetic variations in COVID-19, we engaged RT-PCR. Our study primarily focused on detecting mutations in the RdRp, N, and E genes in Saudi patients with SARS-CoV-2, as well as individuals from various national residing in Saudi Arabia. This molecular technique provided valuable insights into the virus’s genetic makeup during infection and recovery. In our analysis of 671 positive COVID-19 cases, diverse gene involvement patterns were observed. Specifically, 55.91% had mutations in all three genes (RdRp, N, and E), 62.33% in both N and E genes, and 67.16% in RdRp and N genes. Additionally, 30.75% exhibited mutations exclusively in the RdRp gene, and 51.58% had mutations in the N gene. The N gene, in particular, showed high sensitivity as a marker for identifying active viral circulation. Regarding the temporal dynamics of the disease, the median duration between a positive and a subsequent negative COVID-19 RT-PCR test result was approximately 33.86 days for 44% of cases, 14.31 days for 30%, and 22.67 days for 4%. The insights from this study hold significant implications for managing COVID-19 patients during the ongoing pandemic. The N gene shows promise as a marker for detecting active viral circulation, potentially improving patient care and containment strategies. Establishing a defined positive threshold for diagnostic methods and correlating it with a low risk of infection remains a challenge. Further research is needed to address these complexities and enhance our understanding of COVID-19 epidemiology and diagnostics.
Full article
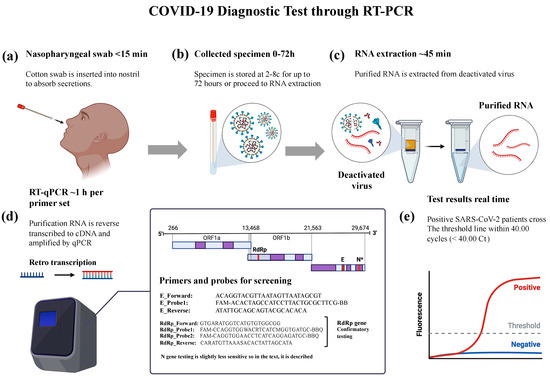
Figure 1
Open AccessCommunication
Evaluation of Genomic Contamination Detection Tools and Influence of Horizontal Gene Transfer on Their Efficiency through Contamination Simulations at Various Taxonomic Ranks
Appl. Microbiol. 2024, 4(1), 124-132; https://doi.org/10.3390/applmicrobiol4010009 - 10 Jan 2024
Abstract
Genomic contamination remains a pervasive challenge in (meta)genomics, prompting the development of numerous detection tools. Despite the attention that this issue has attracted, a comprehensive comparison of the available tools is absent from the literature. Furthermore, the potential effect of horizontal gene transfer
[...] Read more.
Genomic contamination remains a pervasive challenge in (meta)genomics, prompting the development of numerous detection tools. Despite the attention that this issue has attracted, a comprehensive comparison of the available tools is absent from the literature. Furthermore, the potential effect of horizontal gene transfer on the detection of genomic contamination has been little studied. In this study, we evaluated the efficiency of detection of six widely used contamination detection tools. To this end, we developed a simulation framework using orthologous group inference as a robust basis for the simulation of contamination. Additionally, we implemented a variable mutation rate to simulate horizontal transfer. Our simulations covered six distinct taxonomic ranks, ranging from phylum to species. The evaluation of contamination levels revealed the suboptimal precision of the tools, attributed to significant cases of both over-detection and under-detection, particularly at the genus and species levels. Notably, only so-called “redundant” contamination was reliably estimated. Our findings underscore the necessity of employing a combination of tools, including Kraken2, for accurate contamination level assessment. We also demonstrate that none of the assayed tools confused contamination and horizontal gene transfer. Finally, we release CRACOT, a freely accessible contamination simulation framework, which holds promise in evaluating the efficacy of future algorithms.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
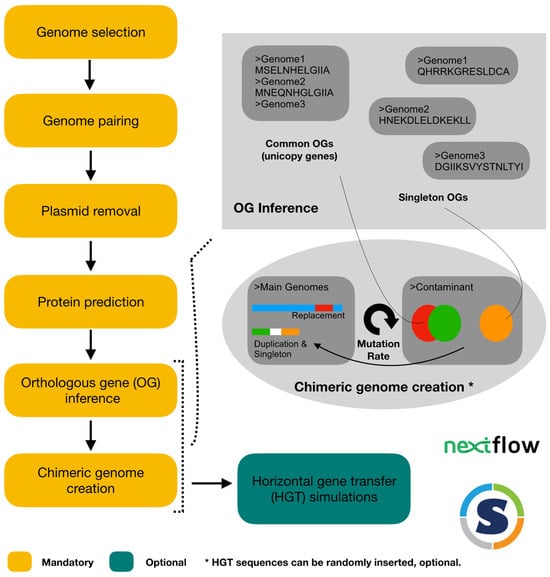
Figure 1
Open AccessArticle
Insights into Genetic and Physiological Characteristics of Clover Rhizobia in Afghanistan Soils
by
, , , , and
Appl. Microbiol. 2024, 4(1), 112-123; https://doi.org/10.3390/applmicrobiol4010008 - 08 Jan 2024
Abstract
Livestock production in Afghanistan highly relies on grazing and clover feed, which is a key component of pastures and forage crops. This study elucidated the genetic diversity of clover-nodulating rhizobia in different ecological regions and their effects on clover growth. A total of
[...] Read more.
Livestock production in Afghanistan highly relies on grazing and clover feed, which is a key component of pastures and forage crops. This study elucidated the genetic diversity of clover-nodulating rhizobia in different ecological regions and their effects on clover growth. A total of 57 rhizobia were isolated and their genetic diversities were studied through 16S rRNA and nifD genes. The isolates were inoculated to clover (Afghan local variety), to investigate the potential of nitrogen fixation and influences of clover growth. The 16S rRNA gene analysis showed two distinct groups of Rhizobium (94.7%) and Ensifer (5.3%) species. The nifD phylogenetic relationship revealed a high similarity to Rhizobium and a novel lineage group close to Rhizobium leguminosarum species. In the plant test, different genotypes significantly (p < 0.01) exhibited an increase in plant biomass production, compared to the un-inoculated plants. Among genotypes, the highest plant biomass was recorded in PC8 (1769.0 mg/plant) and PC9 (1409.2 mg/plant) isolates as compared to un-inoculated plants (144.0 mg/plant). Moreover, these isolates showed maximum nitrogen fixation rates of 8.2 and 6.5 µM/plant, respectively. These isolates were identified as the most promising rhizobial strains for developing biofertilizers in the context of Afghanistan.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
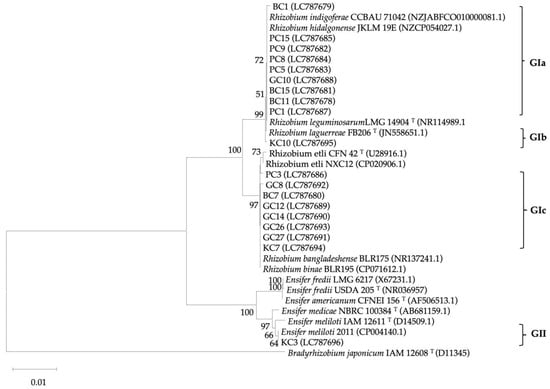
Figure 1
Open AccessArticle
Potential of Lactic Acid Bacteria and Bacillus spp. in a Bio-Detoxification Strategy for Mycotoxin Contaminated Wheat Grains
by
, , , , and
Appl. Microbiol. 2024, 4(1), 96-111; https://doi.org/10.3390/applmicrobiol4010007 - 02 Jan 2024
Abstract
►▼
Show Figures
Mycotoxins present in cereals are a worldwide problem and are a result of the presence of mycotoxin producing fungi. A strategy to reduce these fungi and mycotoxin levels in contaminated grains is with the use of lactic acid bacteria (LAB) or Bacillus spp.,
[...] Read more.
Mycotoxins present in cereals are a worldwide problem and are a result of the presence of mycotoxin producing fungi. A strategy to reduce these fungi and mycotoxin levels in contaminated grains is with the use of lactic acid bacteria (LAB) or Bacillus spp., which can degrade or bind toxins. In this study, LAB and Bacillus spp. were isolated from mycotoxin contaminated wheat grains and, together with additional plant-derived strains, an antifungal screening against Fusarium graminearum was performed. Furthermore, these strains were screened for their ability to reduce zearalenone (ZEA) and deoxynivalenol (DON). Finally, the mode of action of the most promising microorganisms was investigated by analyzing toxin reduction with viable and dead cells, cell extracts and supernatants. Out of 212 tested strains, 70 showed high antifungal activity and 42 exhibited the ability to detoxify more than 90% ZEA, i.e., Bacillus licheniformis (19), B. megaterium (13), and Levilactobacillus brevis (10). None of the tested strains were able to decrease DON. The mode of action of ZEA reduction could not be fully elucidated. Neither dead cells (<20%), nor cell extracts nor supernatants could reduce ZEA in high amounts, which exclude high binding capacity and the involvement of extra- or intra-cellular enzymes.
Full article

Figure 1
Open AccessArticle
Growth and Metabolism of Clostridioides difficile in Hungate-Style Media
Appl. Microbiol. 2024, 4(1), 85-95; https://doi.org/10.3390/applmicrobiol4010006 - 30 Dec 2023
Abstract
►▼
Show Figures
Clostridioides difficile is a clinically and agriculturally important organism with diverse metabolic capabilities. Commercially available media types to cultivate C. difficile typically include multiple growth substrates and often selective agents. Under these conditions, it is difficult to determine what the bacteria utilized and
[...] Read more.
Clostridioides difficile is a clinically and agriculturally important organism with diverse metabolic capabilities. Commercially available media types to cultivate C. difficile typically include multiple growth substrates and often selective agents. Under these conditions, it is difficult to determine what the bacteria utilized and which products are derived from which substrates. These experiments compared a commercial broth (Reinforced Clostridium Medium/RCM) to simpler, defined, carbonate-based media types influenced by Robert Hungate. Peptides (tryptone peptone), amino acids (casamino acids), and/or glucose were added to evaluate the growth of C. difficile strains 9689, BAA-1870, and 43597, and the metabolism of the type strain 9689. C. difficile grew to the greatest optical density in the rich RCM broth but produced less ammonia than the tryptone-containing media types. C. difficile utilized all glucose in RCM and T+G media in addition to performing amino acid fermentations, though the volatile fatty acids produced were not necessarily consistent across media type. When cultured in CAA-containing media, 9689 performed very little metabolism and did not grow regardless of supplementation with glucose. These data demonstrated that C. difficile could metabolize substrates and grow in defined, anaerobic, and carbonate-buffered media. Hungate-style media appear to be an acceptable choice for reliable culturing of C. difficile.
Full article

Figure 1
Open AccessArticle
Occurrence of Mobile Colistin Resistance Genes mcr-1–mcr-10 including Novel mcr Gene Variants in Different Pathotypes of Porcine Escherichia coli Isolates Collected in Germany from 2000 to 2021
by
, , , , and
Appl. Microbiol. 2024, 4(1), 70-84; https://doi.org/10.3390/applmicrobiol4010005 - 28 Dec 2023
Abstract
In the European Union, gastrointestinal disease in pigs is the main indication for the use of colistin, but large-scale epidemiologic data concerning the frequency of mobile colistin resistance (mcr) genes in pig-associated pathotypes of Escherichia coli (E. coli) are
[...] Read more.
In the European Union, gastrointestinal disease in pigs is the main indication for the use of colistin, but large-scale epidemiologic data concerning the frequency of mobile colistin resistance (mcr) genes in pig-associated pathotypes of Escherichia coli (E. coli) are lacking. Multiplex polymerase chain reactions were used to detect virulence-associated genes (VAGs) and mcr-1–mcr-10 genes in 10,573 porcine E. coli isolates collected in Germany from July 2000 to December 2021. Whole genome sequencing was performed on 220 representative mcr-positive E. coli strains. The total frequency of mcr genes was 10.2%, the most frequent being mcr-1 (8.4%) and mcr-4 (1.6%). All other mcr genes were rarely identified (mcr-2, mcr-3, mcr-5) or absent (mcr-6 to mcr-10). The highest frequencies of mcr genes were found in enterotoxigenic and shiga toxin-encoding E. coli (ETEC/STEC hybrid) and in edema disease E. coli (EDEC) strains (21.9% and 17.7%, respectively). We report three novel mcr variants, mcr-1.36, mcr-4.8, and mcr-5.5. In 39 attaching and effacing E. coli (AEEC) isolates analyzed in our study, the eae subtype β1 was the most prevalent (71.8%). Constant surveillance for the presence of mcr genes in various sectors should consider the different frequency of mcr-positive isolates in pathogenic E. coli.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
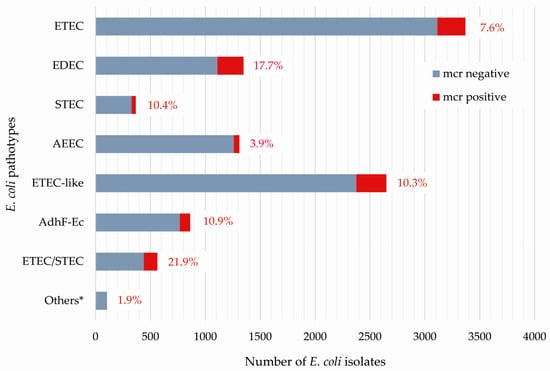
Figure 1
Open AccessReview
Systematic Review of Probiotics and Their Potential for Developing Functional Nondairy Foods
by
and
Appl. Microbiol. 2024, 4(1), 47-69; https://doi.org/10.3390/applmicrobiol4010004 - 26 Dec 2023
Abstract
The gastrointestinal tract is an environment that hosts various microorganisms, including pathogens. Generally, pathogenic bacteria enter the host body through food and the gastrointestinal tract. These pathogenic bacteria can colonize or infiltrate host cells and tissues, causing various infectious diseases. In recent years,
[...] Read more.
The gastrointestinal tract is an environment that hosts various microorganisms, including pathogens. Generally, pathogenic bacteria enter the host body through food and the gastrointestinal tract. These pathogenic bacteria can colonize or infiltrate host cells and tissues, causing various infectious diseases. In recent years, the protective role of probiotic bacteria against gastrointestinal pathogens has been carefully investigated. Probiotics have been found to modulate intestinal microbial flora and play a significant role in the gastrointestinal tract’s function, especially by inhibiting the growth of pathogenic bacteria. However, the mechanism of action of probiotics has yet to be sufficiently proven and recognized. Several important mechanisms support the antagonistic effects of probiotics on various microorganisms, which is achieved, for example, through the production of different antimicrobial compounds, such as bacteriocins, various organic acids, antibiotics, antimicrobial proteins, and exopolysaccharides; mucosal barriers with mucosa and bacteria binding blockers; competition for nutrient uptake; and strengthening of the immune system. Accordingly, this review summarizes the recent studies that have examined the mechanism of action of probiotic bacteria and their beneficial effects in preventing pathogenic bacterial growth and improving gastrointestinal functions. Comprehending their mechanisms of action allows the selection of appropriate probiotic strains for specific applications in gastrointestinal dysfunction.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
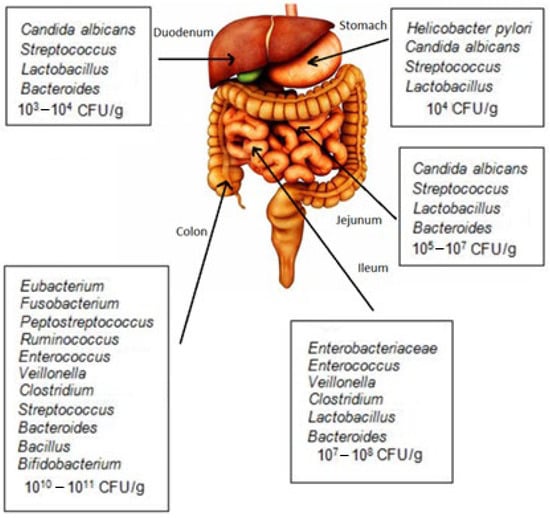
Figure 1
Open AccessArticle
Customizing Sanitization Protocols for Food-Borne Pathogens Based on Biofilm Formation, Surfaces and Disinfectants—Their Two- and Three-Way Interactions
by
, , , , , , and
Appl. Microbiol. 2024, 4(1), 27-46; https://doi.org/10.3390/applmicrobiol4010003 - 23 Dec 2023
Abstract
Food-borne pathogens are a serious challenge in food handling, processing, and packaging systems. The growth of microbial biofilms on food handling surfaces further complicates the management of the microbial contamination of food. Microorganisms within biofilms are difficult to eradicate with chemical disinfectants, with
[...] Read more.
Food-borne pathogens are a serious challenge in food handling, processing, and packaging systems. The growth of microbial biofilms on food handling surfaces further complicates the management of the microbial contamination of food. Microorganisms within biofilms are difficult to eradicate with chemical disinfectants, with an increased likelihood of survival and the subsequent contamination of food. Therefore, a biofilm approach is needed in food safety and hygiene studies. Since many factors, such as strain, cell density, surface type and texture, environmental stress, and so forth, can affect biofilm formation and disinfectant efficacy, we evaluated the responses of biofilms formed by three food-borne bacterial pathogens on eight hard surfaces to seven chemical disinfectants. The three bacteria showed different capacities to colonize the surfaces. Similarly, chemical disinfectants also varied in efficacy, on surfaces and with pathogen species. One-, two-, and three-way interactions of strain, surface, and disinfectant were observed. The results generated demonstrate that the fine-tuning of sanitization strategies along the food production, processing, and packaging chain can be achieved in specific scenarios by accounting for two- and three-way interactions among bacteria, surface, and disinfectant.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
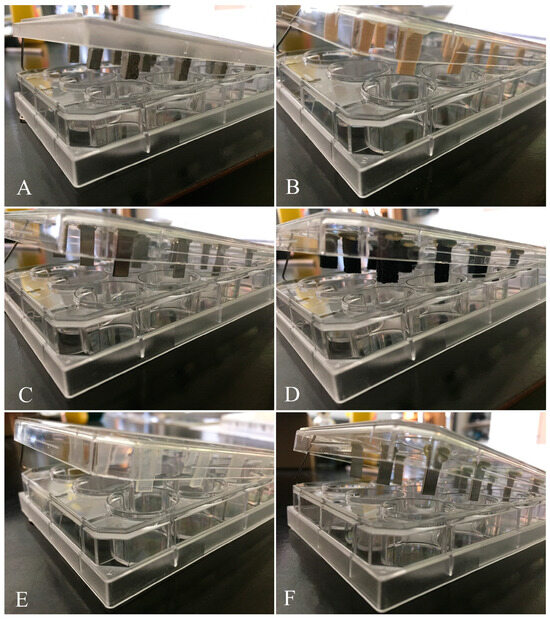

Figure 1
Open AccessArticle
Antimicrobial Resistance Profile of Planctomycetota Isolated from Oyster Shell Biofilm: Ecological Relevance within the One Health Concept
Appl. Microbiol. 2024, 4(1), 16-26; https://doi.org/10.3390/applmicrobiol4010002 - 20 Dec 2023
Abstract
Background: Planctomycetota isolation in pure culture is still challenging with most of the reported data coming from molecular-based methods. Here, we intended to isolate Planctomycetota from the filter-feeder Pacific oyster Magallana gigas, extending the search to a not yet explored natural reservoir
[...] Read more.
Background: Planctomycetota isolation in pure culture is still challenging with most of the reported data coming from molecular-based methods. Here, we intended to isolate Planctomycetota from the filter-feeder Pacific oyster Magallana gigas, extending the search to a not yet explored natural reservoir and to characterize their antimicrobial resistance phenotype. Methods: Oyster samples from different supermarkets and from a farm producer were subject to isolation in selective medium. Inoculation was performed from the shell biofilm and after an enrichment of the edible content. Results: Planctomycetota isolates (n = 65) were only obtained from the shell biofilm with four different species identified: Rhodopirellula baltica (n = 62), Rhodopirellula rubra (n = 1), Rhodopirellula heiligendammensis (n = 1) and Gimesia chilikensis (n = 1). This study reports the first association of Planctomycetota members with oysters and the first description of R. heiligendammensis in Portugal. Moreover, R. rubra, originally identified in Portugal, was isolated from oysters of French origin. Antibiotic susceptibility testing, conducted in strains belonging to two species never assayed before revealed multidrug resistance phenotypes with bacteria showing resistance to several classes of clinically relevant antibiotics (e.g., β-lactams and aminoglycosides). Conclusion: The ecological role and impact of Planctomycetota on oyster holobiont and, ultimately, in public health, under the One Health concept, is discussed.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
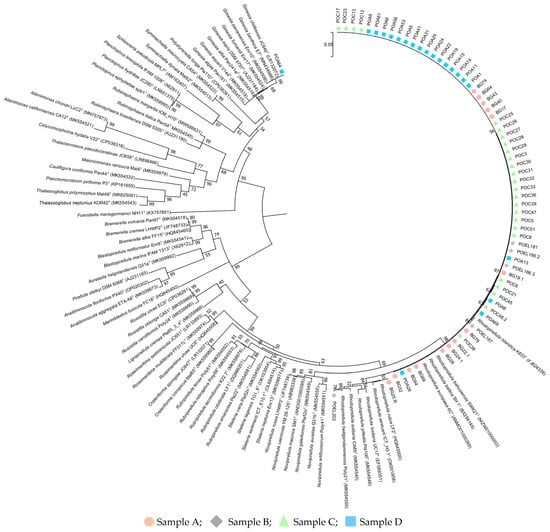
Figure 1
Open AccessReview
Advances in Pretreatment Methods for Free Nucleic Acid Removal in Wastewater Samples: Enhancing Accuracy in Pathogenic Detection and Future Directions
Appl. Microbiol. 2024, 4(1), 1-15; https://doi.org/10.3390/applmicrobiol4010001 - 19 Dec 2023
Abstract
►▼
Show Figures
Accurate pathogenic detection in wastewater is critical for safeguarding public health and the environment. However, the presence of free nucleic acids in wastewater samples poses significant challenges to molecular detection accuracy. This comprehensive review explores the current status and future potential of pretreatment
[...] Read more.
Accurate pathogenic detection in wastewater is critical for safeguarding public health and the environment. However, the presence of free nucleic acids in wastewater samples poses significant challenges to molecular detection accuracy. This comprehensive review explores the current status and future potential of pretreatment methods to remove free nucleic acids from wastewater samples. The study contributes a comprehensive analysis of the mechanisms, strengths, and limitations of various pretreatment approaches, including physical, chemical, and enzymatic processes. The effect of various factors on the removal efficiency of these pretreatment methods is also discussed. This review enhances our comprehension of pretreatment techniques and their vital role in achieving precise pathogenic detection in complex wastewater matrices. Furthermore, it outlines future perspectives and developments for improving the speed and effectiveness of pathogenic detection, contributing significantly to disease surveillance, early warning systems, and environmental protection.
Full article

Figure 1
Open AccessArticle
The Impact of Lytic Viruses on Bacterial Carbon Metabolism in a Temperate Freshwater Reservoir (Naussac, France)
by
, , , , and
Appl. Microbiol. 2023, 3(4), 1407-1423; https://doi.org/10.3390/applmicrobiol3040095 - 15 Dec 2023
Abstract
In aquatic systems, the impact of the viral regulation of bacterial carbon metabolism (BCM) is often overlooked compared with nutrient supply. To address this gap, an investigation was conducted in the euphotic and aphotic zones of a mesotrophic freshwater reservoir (Naussac, France) to
[...] Read more.
In aquatic systems, the impact of the viral regulation of bacterial carbon metabolism (BCM) is often overlooked compared with nutrient supply. To address this gap, an investigation was conducted in the euphotic and aphotic zones of a mesotrophic freshwater reservoir (Naussac, France) to assess the relative influence of lytic viral infection on key bacterial metabolic parameters, specifically bacterial production (BP) and respiration (BR), as indicators of BCM. Measured using flow cytometry, the abundance of viral sub-groups (V1–V3) exhibited a consistent pattern in tandem with their bacterial hosts across both time and space. A more significant relationship between bacterial and viral parameters than between physicochemical factors suggested a prevailing internal control mechanism that was potentially driven by viral lysis. Viral-mediated bacterial mortality up to 65% was evident in the euphotic zone. The observed variation in BCM (ranging from 7% to 32%) was explained by an uncoupling between BR and BP. Notably, BR was significantly higher (three-fold) than BP in bacterial communities subjected to low in situ phosphate concentrations (<0.5 µM P) and high nutrient stoichiometric ratios (N:P > 60, C:P > 900). An antagonistic relationship between lytic viruses and BCM, whereby the repression of bacterial growth results in elevated respiratory demands, could potentially be attributed to substrate availability constraints.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
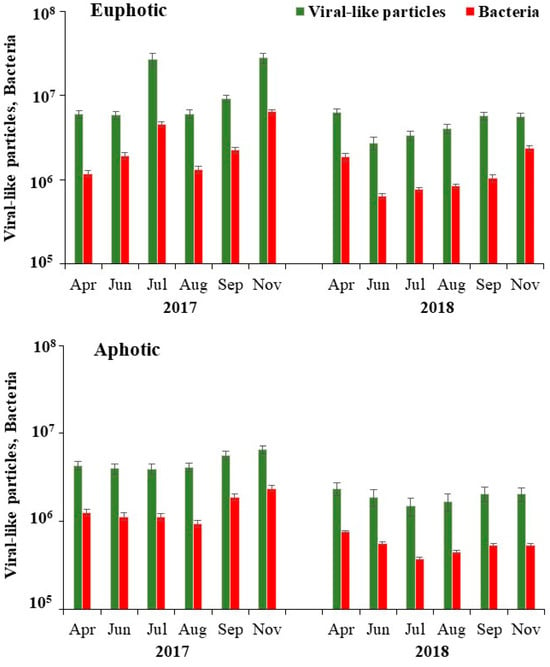
Figure 1
Open AccessArticle
Campylobacter Bacteriophage Infection at Refrigeration Temperatures
Appl. Microbiol. 2023, 3(4), 1392-1406; https://doi.org/10.3390/applmicrobiol3040094 - 13 Dec 2023
Abstract
The application of bacteriophages to control foodborne bacterial pathogens in foods has gained traction in recent years. Poultry meat is a major source of Campylobacter jejuni, and a target for the application of bacteriophages. To offer the prospect of a post-harvest control
[...] Read more.
The application of bacteriophages to control foodborne bacterial pathogens in foods has gained traction in recent years. Poultry meat is a major source of Campylobacter jejuni, and a target for the application of bacteriophages. To offer the prospect of a post-harvest control measure, the bacteriophage must function at refrigeration temperatures, where C. jejuni does not grow but can survive. Here, we report actions of three classes of Campylobacter bacteriophage at 4 °C. The pre-incubation of broth cultures at 4 °C before a shift to 42 °C under conditions that support the growth of the host bacteria revealed differences in the time to lysis compared with cultures incubated at 42 °C. The pre-adsorption of the bacteriophage to a sub-population of bacteria is consistent with the observation of asynchronous infection. To ascertain whether the bacteriophages adsorb and infect (the commitment to replicate), we investigated bacteriophage transcription at 4 °C. RNA transcripts for all the bacteriophage host combinations were detected after 15 min, indicating that the interaction is not merely passive. Bacteriophages can infect C. jejuni at refrigeration temperatures, but the infection does not proceed to lysis in the absence of host cell division.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
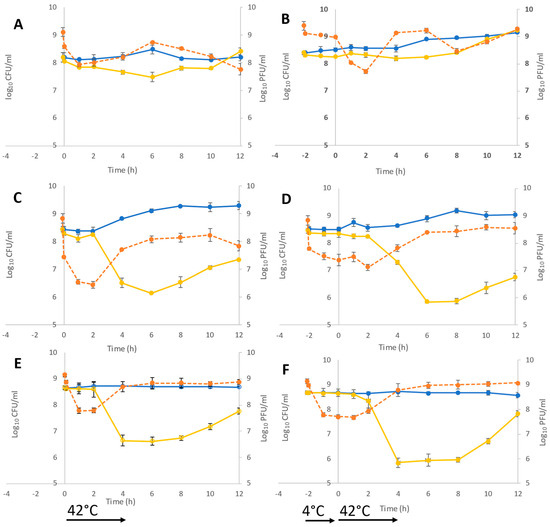
Figure 1
Open AccessEditorial
Antimicrobial Resistance in the Time of COVID-19
Appl. Microbiol. 2023, 3(4), 1388-1391; https://doi.org/10.3390/applmicrobiol3040093 - 11 Dec 2023
Abstract
The world is presently dealing with two pandemics—COVID-19 and antibiotic resistance (AMR)—that constitute a serious menace to public health on a worldwide basis [...]
Full article
(This article belongs to the Topic Antimicrobial Resistance in the Time of COVID-19)
Open AccessArticle
The Impact of Formulation and Freeze Drying on the Properties and Performance of Freeze-Dried Limosilactobacillus reuteri R2LC
by
, , , , , , and
Appl. Microbiol. 2023, 3(4), 1370-1387; https://doi.org/10.3390/applmicrobiol3040092 - 03 Dec 2023
Abstract
►▼
Show Figures
Freeze drying is a commonly used method for preserving probiotic bacteria and live biotherapeutic products. Before drying, the bacterial cells are formulated with a lyoprotectant, and the design of these two process steps are crucial to achieve a high-quality product. There are several
[...] Read more.
Freeze drying is a commonly used method for preserving probiotic bacteria and live biotherapeutic products. Before drying, the bacterial cells are formulated with a lyoprotectant, and the design of these two process steps are crucial to achieve a high-quality product. There are several factors that may affect the biological and physicochemical properties of the freeze-dried cells and we have used a Design of Experiment approach to investigate the effects of formulation and freeze-drying parameters on properties and performance of Limosilactobacillus reuteri R2LC. The biological characteristics of the dried bacteria were evaluated by measuring cell survival, metabolic activity and stability, and physicochemical characteristics were studied using visual inspection, differential scanning calorimetry (DSC), scanning electron microscopy (SEM), and analysis of residual moisture content and bacterial aggregation. A comparison between the lyoprotectants trehalose and sucrose showed that the latter gave better freeze-drying survival, metabolic activity, and storage stability. We also want to highlight that there was a correlation between bacterial concentration, metabolic activity, and aggregation of bacteria, where a higher concentration (1010 CFU/mL) resulted in both higher metabolic activity and aggregation. Several other process and formulation factors affected both the biological and physicochemical properties of freeze-dried L. reuteri R2LC and it could be concluded that care must be taken to develop a production method that generates a product with high and consistent quality. These results may, or may not, be strain specific.
Full article

Graphical abstract
Open AccessArticle
Employing Active Learning in Medium Optimization for Selective Bacterial Growth
Appl. Microbiol. 2023, 3(4), 1355-1369; https://doi.org/10.3390/applmicrobiol3040091 - 03 Dec 2023
Abstract
Medium optimization and development for selective bacterial cultures are essential for isolating and functionalizing individual bacteria in microbial communities; nevertheless, it remains challenging due to the unknown mechanisms between bacterial growth and medium components. The present study first tried combining machine learning (ML)
[...] Read more.
Medium optimization and development for selective bacterial cultures are essential for isolating and functionalizing individual bacteria in microbial communities; nevertheless, it remains challenging due to the unknown mechanisms between bacterial growth and medium components. The present study first tried combining machine learning (ML) with active learning to fine-tune the medium components for the selective culture of two divergent bacteria, i.e., Lactobacillus plantarum and Escherichia coli. ML models considering multiple growth parameters of the two bacterial strains were constructed to predict the fine-tuned medium combinations for higher specificity of bacterial growth. The growth parameters were designed as the exponential growth rate (r) and maximal growth yield (K), which were calculated according to the growth curves. The eleven chemical components in the commercially available medium MRS were subjected to medium optimization and specialization. High-throughput growth assays of both strains grown separately were performed to obtain thousands of growth curves in more than one hundred medium combinations, and the resultant datasets linking the growth parameters to the medium combinations were used for the ML training. Repeated rounds of active learning (i.e., ML model construction, medium prediction, and experimental verification) successfully improved the specific growth of a single strain out of the two. Both r and K showed maximized differentiation between the two strains. A further analysis of all the data accumulated in active learning identified the decision-making medium components for growth specificity and the differentiated, determinative manner of growth decisions of the two strains. In summary, this study demonstrated the efficiency and practicality of active learning in medium optimization for selective cultures and offered novel insights into the contribution of the chemical components to specific bacterial growth.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
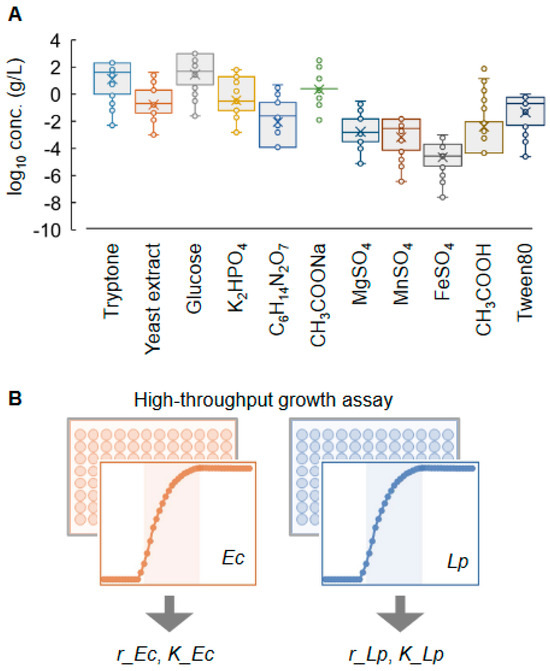
Figure 1
Open AccessArticle
Reduced Infestation Levels of Lepeophtheirus salmonis in Atlantic Salmon (Salmo salar) following Immersion Exposure to Probiotic Aliivibrio spp.
by
, , , , , and
Appl. Microbiol. 2023, 3(4), 1339-1354; https://doi.org/10.3390/applmicrobiol3040090 - 30 Nov 2023
Abstract
Salmon lice (Lepeophtheirus salmonis) constitute a major challenge during the production of farmed Atlantic salmon in Norway. Preventive measures are considered to have a higher impact on sustainable control than lice treatment. Therefore, the studies presented here aimed to document the
[...] Read more.
Salmon lice (Lepeophtheirus salmonis) constitute a major challenge during the production of farmed Atlantic salmon in Norway. Preventive measures are considered to have a higher impact on sustainable control than lice treatment. Therefore, the studies presented here aimed to document the preventive effects of probiotic Aliivibrio spp. on lice infestation in experimental challenges. A reduction in salmon lice attachment success (58–65%) was observed in two separate aquarium trials, where Atlantic salmon were exposed to different compositions of Aliivibrio species 91 and 155 days prior to lice challenge. In a third trial, no difference in attachment was observed in groups exposed to probiotics 58 days prior to lice challenge compared to controls. However, a relative reduction in lice counts was seen on movable stages later in the trial. High levels of probiotic bacteria had no impact on lice viability in an in vitro bioassay on the preadult life stage; thus, the mechanism behind the preventative effect remains unknown. In conclusion, probiotic Aliivibrio bacteria can likely be used as a preventive tool to reduce salmon louse infestations in the salmon industry. The mechanism is still unknown, and this novel approach to lice control warrants further investigation to understand its optimal use and potential.
Full article
(This article belongs to the Special Issue Current Trends in the Applications of Probiotics and Other Beneficial Microbes)
►▼
Show Figures

Figure 1
Open AccessReview
The Vaginal Microbiome during Pregnancy in Health and Disease
Appl. Microbiol. 2023, 3(4), 1302-1338; https://doi.org/10.3390/applmicrobiol3040089 - 30 Nov 2023
Abstract
This study appraises the progress in the understanding of the composition of the vaginal microflora with a focus on the microbiome during pregnancy. This knowledge is presented with the background of the global health contribution, along with the importance of these microbial communities
[...] Read more.
This study appraises the progress in the understanding of the composition of the vaginal microflora with a focus on the microbiome during pregnancy. This knowledge is presented with the background of the global health contribution, along with the importance of these microbial communities to pregnancy. A brief review of current methods employed to investigate the structure of these microbial populations is included. Two types of studies, cross-sectional and longitudinal, have been used to characterise the vaginal microbiota; both types are reviewed since they provide information that serves to piece together a more complete picture of the vaginal microflora and its changes during pregnancy. The identity of microbes present in the vagina are examined in the context of health and disease, and, more specifically, in the setting of pregnancy outcomes. The protective role of lactobacilli in maintaining a healthy vaginal environment is evaluated, with analyses of the different roles of various Lactobacillus spp. Classifications of the vaginal microbiota into vagitypes in non-pregnant and pregnant women are discussed. The associations of specific taxa with three adverse pregnancy results, namely, miscarriage, stillbirth, and preterm birth, are examined in some detail. Longitudinal studies investigating changes in the bacterial community composition and taxa abundance demonstrate that this microbiota decreases in richness and diversity relative to those present in non-pregnant microbiomes. Notwithstanding the significant effort made to characterise the vagina bacterial microbiota, a large number of issues remain to be fully understood.
Full article
(This article belongs to the Special Issue Microbiome in Ecosystem 2.0)
►▼
Show Figures
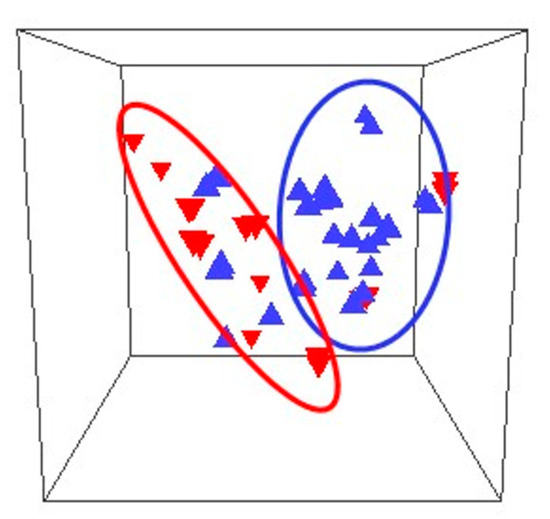
Figure 1
Open AccessArticle
Assessment of the Spoilage Microbiota and the Growth Potential of Listeria monocytogenes in Minced Free-Range Chicken Meat Stored at 4 °C in Vacuum: Comparison with the Spoilage Community of Resultant Retail Modified Atmosphere Packaged Products
Appl. Microbiol. 2023, 3(4), 1277-1301; https://doi.org/10.3390/applmicrobiol3040088 - 28 Nov 2023
Abstract
Although current diet and nutrition trends in developed countries led the poultry industry to shift to alternative breeding/production methods, such as organic and free-range, limited data on the microbiology of alternative compared to conventional poultry meat products exist. Therefore, this study assessed the
[...] Read more.
Although current diet and nutrition trends in developed countries led the poultry industry to shift to alternative breeding/production methods, such as organic and free-range, limited data on the microbiology of alternative compared to conventional poultry meat products exist. Therefore, this study assessed the evolution and composition of the spoilage microbiota and the growth potential of inoculated (3 log cfu/g) Listeria monocytogenes in freshly minced free-range chicken meat stored at 4 °C in vacuum packages (VP; four batches) for 0, 3, 5, 7, and 10 days. Additionally, two VP batches were compared with their resultant retail products stored in modified atmosphere packages (MAP 30:70 CO2/N2) at 4 °C to detect potential differences with the MAP spoilage community described previously. The initial pH of the VP minces was 6.0–6.1, except for one mince, designated VP + AA, which had initial pH 5.8 and was found to contain ‘external’ 1.26% L-lactate and 0.24% acetate associated with a vinegar smell during storage. The rest of the VP batches contained on average 0.75% L-lactate and 0.02% acetate on day 0. After 7 days at 4 °C, L-lactate decreased by at least 3-fold in VP and over 5-fold in VP + AA vs. minor decreases in MAP. Acetate increased 2-fold in all batches. D-lactate (ca. 0.02% on day 0) increased by 4-fold in VP batches only. Lactic acid bacteria (LAB) became the dominant spoilers in all samples. Only VP allowed a delayed 10-fold growth (>5.0 to 6.2 log cfu/g) of pseudomonads from day 7 to day 10 at 4 °C. Compared to VP, VP + AA and MAP retarded growth of LAB, pseudomonads, and enterobacteria by 1–2 log units, at final levels below 6.5, 4.5, and 3.0 log cfu/g, respectively. Enterococci, staphylococci, yeasts, and L. monocytogenes did not grow. Latilactobacillus sakei predominated in all spoiled VP batches (65.8% of 80 meat isolates) followed by Latilactobacillus fuchuensis (9.2%), Leuconostoc carnosum (6.6%), Carnobacterium divergens (6.6%), Latilactobacillus curvatus (5.3%), and Weissella koreensis (2.6%). VP + AA favored Latilactobacillus. Brochothrix thermosphacta was frequent in one VP batch. In conclusion, cold-stored (4 °C), minced, free-range chicken meat spoils more rapidly and offensively under VP than MAP or VP combined with acetate-containing (VP + AA) antimicrobial blends.
Full article
(This article belongs to the Special Issue Applied Microbiology of Foods 2.0)
►▼
Show Figures
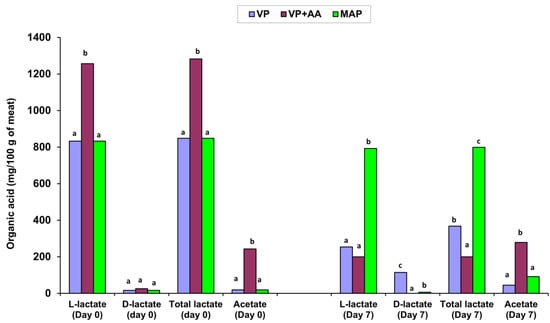
Figure 1
Open AccessArticle
Soil Bacteriome Resilience and Reduced Nitrogen Toxicity in Tomato by Controlled Release Nitrogen Fertilizer Compared to Urea
Appl. Microbiol. 2023, 3(4), 1262-1276; https://doi.org/10.3390/applmicrobiol3040087 - 20 Nov 2023
Abstract
Controlled release fertilizers (CRFs) mitigate negative effects of high nitrogen (N) fertilization rates, such as N toxicity and soil N loss. However, it is unknown if potentially toxic rates of CRF and quick release fertilizer differentially affect soil bacterial communities. To examine potential
[...] Read more.
Controlled release fertilizers (CRFs) mitigate negative effects of high nitrogen (N) fertilization rates, such as N toxicity and soil N loss. However, it is unknown if potentially toxic rates of CRF and quick release fertilizer differentially affect soil bacterial communities. To examine potential N toxicity effects on soil microbial communities, we grew tomato (Solanum lycopersicum “Rutgers”) for eight weeks in soils that were fertilized with high levels of quick release or controlled release urea and in soils with either low or high initial microbial N competitor populations. In both soils, we observed N toxicity in urea-fertilized tomatoes, but toxicity was ameliorated with CRF application. Controlled release fertilization increased soil N retention, thereby reducing soil N loss. While N toxicity symptoms manifested in the plant, the soil microbiome was only minorly affected. There were subtle differences in soil bacterial populations, in which nitrifying bacteria accumulated in soils fertilized at high N rates, regardless of the type of N fertilizer used. Ultimately, CRF reduced plant N toxicity symptoms but did not change the soil microbiome compared to quick release urea. These results show that while there are clear benefits of CRF regarding N toxicity tolerance on crops, the soil microbiome is resilient to this abiotic stressor.
Full article
(This article belongs to the Special Issue Microbiome in Ecosystem 2.0)
►▼
Show Figures
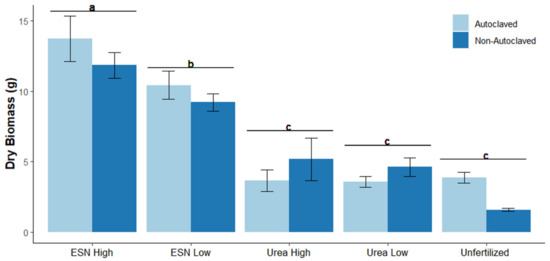
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Microbiology, Bioengineering, Biology, Environments, Microorganisms
Environmental Bioengineering and Geomicrobiology
Topic Editors: Xian-Chun Zeng, Deng LiuDeadline: 20 December 2024

Conferences
Special Issues
Special Issue in
Applied Microbiology
Applied Microbiology of Foods 2.0
Guest Editor: Peter MurianaDeadline: 31 January 2024
Special Issue in
Applied Microbiology
Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024)
Guest Editor: Ian ConnertonDeadline: 30 June 2024
Special Issue in
Applied Microbiology
Microbiome in Ecosystem 3.0
Guest Editor: Bong-Soo KimDeadline: 31 October 2024
Special Issue in
Applied Microbiology
Current Trends in the Applications of Probiotics and Other Beneficial Microbes
Guest Editor: Sabina FijanDeadline: 31 December 2024












