Journal Description
BioMedInformatics
BioMedInformatics
is an international, peer-reviewed, open access journal on all areas of biomedical informatics, as well as computational biology and medicine, published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, EBSCO, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 21 days after submission; acceptance to publication is undertaken in 8.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
Latest Articles
Factors Associated with Unplanned Hospital Readmission after Discharge: A Descriptive and Predictive Study Using Electronic Health Record Data
BioMedInformatics 2024, 4(1), 219-235; https://doi.org/10.3390/biomedinformatics4010014 - 12 Jan 2024
Abstract
Hospital readmission involves the unplanned emergency admission of patients within 30 days from discharge after the previous admission. According to the Canadian Health Institute (CIHI), 1 in 11 patients were readmitted within 30 days of leaving the hospital in 2021. In the USA,
[...] Read more.
Hospital readmission involves the unplanned emergency admission of patients within 30 days from discharge after the previous admission. According to the Canadian Health Institute (CIHI), 1 in 11 patients were readmitted within 30 days of leaving the hospital in 2021. In the USA, nearly 20% of Medicare patients were readmitted after discharge, where the average cost of readmission was approximately USD 15,000, as reported by the Agency for Healthcare Research and Quality (AHQR) in 2018. To tackle this issue, we first conducted a descriptive analysis study to understand the risk factors associated with hospital readmission, and then we applied machine learning approaches to predict hospital readmission by using patients’ demographic and clinical data extracted from the Electronic Health Record of the MIMIC-III clinical database. The results showed that the number of previous admissions during the last 12 months, hyperosmolar imbalance and comorbidity index were the top three significant factors for hospital readmission. The predictive model achieved a performance of 95.6% AP and an AUC = 97.3% using the Gradient Boosting algorithm trained on all features.
Full article
(This article belongs to the Special Issue Feature Papers in Computational Biology and Medicine)
►
Show Figures
Open AccessArticle
An Explainable AI System for the Diagnosis of High-Dimensional Biomedical Data
by
, , , , , and
BioMedInformatics 2024, 4(1), 197-218; https://doi.org/10.3390/biomedinformatics4010013 - 11 Jan 2024
Abstract
Typical state-of-the-art flow cytometry data samples typically consist of measures of 10 to 30 features of more than 100,000 cell “events”. Artificial intelligence (AI) systems are able to diagnose such data with almost the same accuracy as human experts. However, such systems face
[...] Read more.
Typical state-of-the-art flow cytometry data samples typically consist of measures of 10 to 30 features of more than 100,000 cell “events”. Artificial intelligence (AI) systems are able to diagnose such data with almost the same accuracy as human experts. However, such systems face one central challenge: their decisions have far-reaching consequences for the health and lives of people. Therefore, the decisions of AI systems need to be understandable and justifiable by humans. In this work, we present a novel explainable AI (XAI) method called algorithmic population descriptions (ALPODS), which is able to classify (diagnose) cases based on subpopulations in high-dimensional data. ALPODS is able to explain its decisions in a form that is understandable to human experts. For the identified subpopulations, fuzzy reasoning rules expressed in the typical language of domain experts are generated. A visualization method based on these rules allows human experts to understand the reasoning used by the AI system. A comparison with a selection of state-of-the-art XAI systems shows that ALPODS operates efficiently on known benchmark data and on everyday routine case data.
Full article
(This article belongs to the Section Computational Biology and Medicine)
►▼
Show Figures

Graphical abstract
Open AccessReview
Digital Pathology: A Comprehensive Review of Open-Source Histological Segmentation Software
by
, , , , , , , and
BioMedInformatics 2024, 4(1), 173-196; https://doi.org/10.3390/biomedinformatics4010012 - 11 Jan 2024
Abstract
In the era of digitalization, the biomedical sector has been affected by the spread of artificial intelligence. In recent years, the possibility of using deep and machine learning methods for clinical diagnostic and therapeutic interventions has been emerging as an essential resource for
[...] Read more.
In the era of digitalization, the biomedical sector has been affected by the spread of artificial intelligence. In recent years, the possibility of using deep and machine learning methods for clinical diagnostic and therapeutic interventions has been emerging as an essential resource for biomedical imaging. Digital pathology represents innovation in a clinical world that looks for faster and better-performing diagnostic methods, without losing the accuracy of current human-guided analyses. Indeed, artificial intelligence has played a key role in a wide variety of applications that require the analysis of a massive amount of data, including segmentation processes in medical imaging. In this context, artificial intelligence enables the improvement of image segmentation methods, moving towards the development of fully automated systems of analysis able to support pathologists in decision-making procedures. The aim of this review is to aid biologists and clinicians in discovering the most common segmentation open-source tools, including ImageJ (v. 1.54), CellProfiler (v. 4.2.5), Ilastik (v. 1.3.3) and QuPath (v. 0.4.3), along with their customized implementations. Additionally, the tools’ role in the histological imaging field is explored further, suggesting potential application workflows. In conclusion, this review encompasses an examination of the most commonly segmented tissues and their analysis through open-source deep and machine learning tools.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
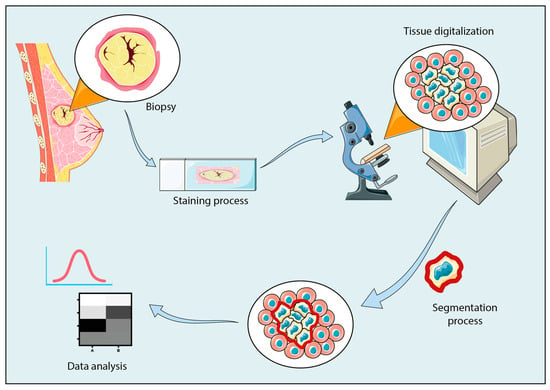
Figure 1
Open AccessReview
Small Bowel Dose Constraints in Radiation Therapy—Where Omics-Driven Biomarkers and Bioinformatics Can Take Us in the Future
BioMedInformatics 2024, 4(1), 158-172; https://doi.org/10.3390/biomedinformatics4010011 - 11 Jan 2024
Abstract
Radiation-induced gastrointestinal (GI) dose constraints are still a matter of concern with the ongoing evolution of patient outcomes and treatment-related toxicity in the era of image-guided intensity-modulated radiation therapy (IMRT), stereotactic ablative radiotherapy (SABR), and novel systemic agents. Small bowel (SB) dose constraints
[...] Read more.
Radiation-induced gastrointestinal (GI) dose constraints are still a matter of concern with the ongoing evolution of patient outcomes and treatment-related toxicity in the era of image-guided intensity-modulated radiation therapy (IMRT), stereotactic ablative radiotherapy (SABR), and novel systemic agents. Small bowel (SB) dose constraints in pelvic radiotherapy (RT) are a critical aspect of treatment planning, and prospective data to support them are scarce. Previous and current guidelines are based on retrospective data and experts’ opinions. Patient-related factors, including genetic, biological, and clinical features and systemic management, modulate toxicity. Omic and microbiome alterations between patients receiving RT to the SB may aid in the identification of patients at risk and real-time identification of acute and late toxicity. Actionable biomarkers may represent a pragmatic approach to translating findings into personalized treatment with biologically optimized dose escalation, given the mitigation of the understood risk. Biomarkers grounded in the genome, transcriptome, proteome, and microbiome should undergo analysis in trials that employ, R.T. Bioinformatic templates will be needed to help advance data collection, aggregation, and analysis, and eventually, decision making with respect to dose constraints in the modern RT era.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
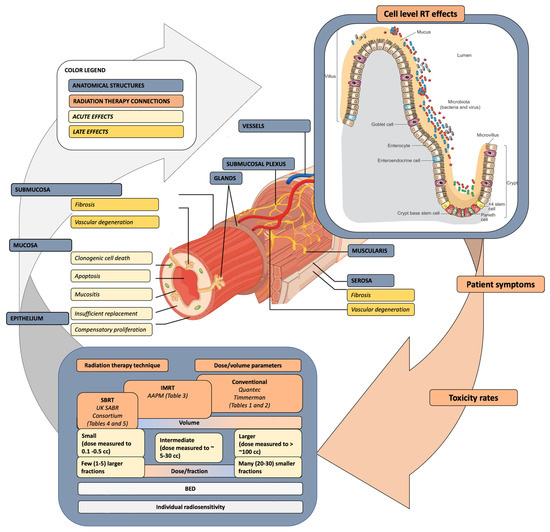
Figure 1
Open AccessArticle
Blood Pressure Estimation from Photoplythmography Using Hybrid Scattering–LSTM Networks
by
, , , , and
BioMedInformatics 2024, 4(1), 139-157; https://doi.org/10.3390/biomedinformatics4010010 - 09 Jan 2024
Abstract
One of the most significant indicators of heart and cardiovascular health is blood pressure (BP). Blood pressure (BP) has gained great attention in the last decade. Uncontrolled high blood pressure increases the risk of serious health problems, including heart attack and stroke. Recently,
[...] Read more.
One of the most significant indicators of heart and cardiovascular health is blood pressure (BP). Blood pressure (BP) has gained great attention in the last decade. Uncontrolled high blood pressure increases the risk of serious health problems, including heart attack and stroke. Recently, machine/deep learning has been leveraged for learning a BP from photoplethysmography (PPG) signals. Hence, continuous BP monitoring can be introduced, based on simple wearable contact sensors or even remotely sensed from a proper camera away from the clinical setup. However, the available training dataset imposes many limitations besides the other difficulties related to the PPG time series as high-dimensional data. This work presents beat-by-beat continuous PPG-based BP monitoring while accounting for the aforementioned limitations. For a better exploration of beats’ features, we propose to use wavelet scattering transform as a better descriptive domain to cope with the limitation of the training dataset and to help the deep learning network accurately learn the relationship between the morphological shapes of PPG beats and the BP. A long short-term memory (LSTM) network is utilized to demonstrate the superiority of the wavelet scattering transform over other domains. The learning scenarios are carried out on a beat basis where the input corresponding PPG beat is used for predicting BP in two scenarios; (1) Beat-by-beat arterial blood pressure (ABP) estimation, and (2) Beat-by-beat estimation of the systolic and diastolic blood pressure values. Different transformations are used to extract the features of the PPG beats in different domains including time, discrete cosine transform (DCT), discrete wavelet transform (DWT), and wavelet scattering transform (WST) domains. The simulation results show that using the WST domain outperforms the other domains in the sense of root mean square error (RMSE) and mean absolute error (MAE) for both of the suggested two scenarios.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
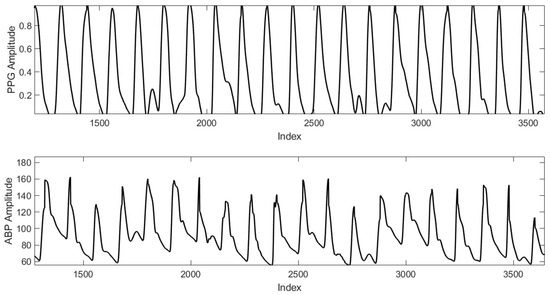
Figure 1
Open AccessArticle
Machine Learning Analysis of Genomic Factors Influencing Hyperbaric Oxygen Therapy in Parkinson’s Disease
BioMedInformatics 2024, 4(1), 127-138; https://doi.org/10.3390/biomedinformatics4010009 - 09 Jan 2024
Abstract
(1) Background: Parkinson’s disease (PD) is a progressively worsening neurodegenerative disorder affecting movement, mental well-being, sleep, and pain. While no cure exists, treatments like hyperbaric oxygen therapy (HBOT) offer potential relief. However, the molecular biology perspective, especially when intertwined with machine learning dynamics,
[...] Read more.
(1) Background: Parkinson’s disease (PD) is a progressively worsening neurodegenerative disorder affecting movement, mental well-being, sleep, and pain. While no cure exists, treatments like hyperbaric oxygen therapy (HBOT) offer potential relief. However, the molecular biology perspective, especially when intertwined with machine learning dynamics, remains underexplored. (2) Methods: We employed machine learning techniques to analyze single-cell RNA-seq data from human PD cell samples. This approach aimed to identify pivotal genes associated with PD and understand their relationship with HBOT. (3) Results: Our analysis indicated genes such as MAP2, CAP2, and WSB1, among others, as being crucially linked with Parkinson’s disease (PD) and showed their significant correlation with Hyperbaric oxygen therapy (HBOT) indicatively. This suggests that certain genomic factors might influence the efficacy of HBOT in PD treatment. (4) Conclusions: HBOT presents promising therapeutic potential for Parkinson’s disease, with certain genomic factors playing a pivotal role in its efficacy. Our findings emphasize the need for further machine learning-driven research harnessing diverse omics data to better understand and treat PD.
Full article
(This article belongs to the Special Issue Feature Papers in Medical Statistics and Data Science Section)
►▼
Show Figures
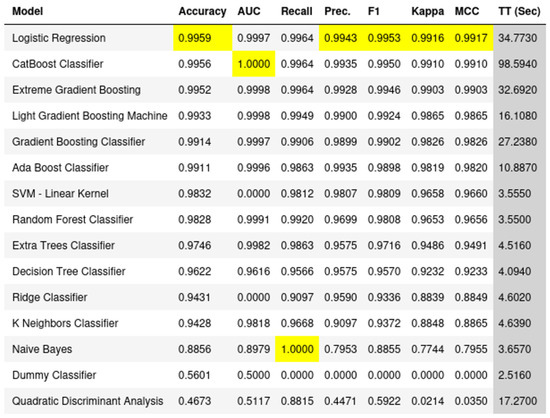
Figure 1
Open AccessReview
Interpretable Medical Imagery Diagnosis with Self-Attentive Transformers: A Review of Explainable AI for Health Care
by
BioMedInformatics 2024, 4(1), 113-126; https://doi.org/10.3390/biomedinformatics4010008 - 08 Jan 2024
Abstract
Recent advancements in artificial intelligence (AI) have facilitated its widespread adoption in primary medical services, addressing the demand–supply imbalance in healthcare. Vision Transformers (ViT) have emerged as state-of-the-art computer vision models, benefiting from self-attention modules. However, compared to traditional machine learning approaches, deep
[...] Read more.
Recent advancements in artificial intelligence (AI) have facilitated its widespread adoption in primary medical services, addressing the demand–supply imbalance in healthcare. Vision Transformers (ViT) have emerged as state-of-the-art computer vision models, benefiting from self-attention modules. However, compared to traditional machine learning approaches, deep learning models are complex and are often treated as a “black box” that can cause uncertainty regarding how they operate. Explainable artificial intelligence (XAI) refers to methods that explain and interpret machine learning models’ inner workings and how they come to decisions, which is especially important in the medical domain to guide healthcare decision-making processes. This review summarizes recent ViT advancements and interpretative approaches to understanding the decision-making process of ViT, enabling transparency in medical diagnosis applications.
Full article
(This article belongs to the Special Issue Explainable Artificial Intelligence (XAI) in Biomedical Research and Clinical Practice)
►▼
Show Figures
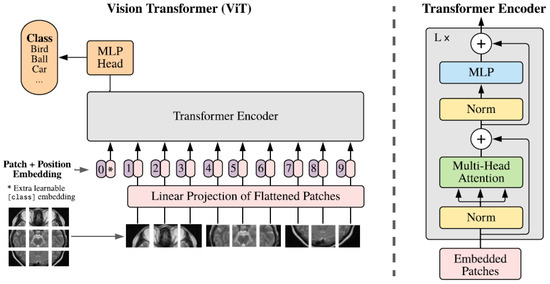
Figure 1
Open AccessArticle
Limitations of Protein Structure Prediction Algorithms in Therapeutic Protein Development
BioMedInformatics 2024, 4(1), 98-112; https://doi.org/10.3390/biomedinformatics4010007 - 08 Jan 2024
Abstract
The three-dimensional protein structure is pivotal in comprehending biological phenomena. It directly governs protein function and hence aids in drug discovery. The development of protein prediction algorithms, such as AlphaFold2, ESMFold, and trRosetta, has given much hope in expediting protein-based therapeutic discovery. Though
[...] Read more.
The three-dimensional protein structure is pivotal in comprehending biological phenomena. It directly governs protein function and hence aids in drug discovery. The development of protein prediction algorithms, such as AlphaFold2, ESMFold, and trRosetta, has given much hope in expediting protein-based therapeutic discovery. Though no study has reported a conclusive application of these algorithms, the efforts continue with much optimism. We intended to test the application of these algorithms in rank-ordering therapeutic proteins for their instability during the pre-translational modification stages, as may be predicted according to the confidence of the structure predicted by these algorithms. The selected molecules were based on a harmonized category of licensed therapeutic proteins; out of the 204 licensed products, 188 that were not conjugated were chosen for analysis, resulting in a lack of correlation between the confidence scores and structural or protein properties. It is crucial to note here that the predictive accuracy of these algorithms is contingent upon the presence of the known structure of the protein in the accessible database. Consequently, our conclusion emphasizes that these algorithms primarily replicate information derived from existing structures. While our findings caution against relying on these algorithms for drug discovery purposes, we acknowledge the need for a nuanced interpretation. Considering their limitations and recognizing that their utility may be constrained to scenarios where known structures are available is important. Hence, caution is advised when applying these algorithms to characterize various attributes of therapeutic proteins without the support of adequate structural information. It is worth noting that the two main algorithms, AlfphaFold2 and ESMFold, also showed a 72% correlation in their scores, pointing to similar limitations. While much progress has been made in computational sciences, the Levinthal paradox remains unsolved.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures

Graphical abstract
Open AccessEditorial
Biomedical Informatics: State of the Art, Challenges, and Opportunities
BioMedInformatics 2024, 4(1), 89-97; https://doi.org/10.3390/biomedinformatics4010006 - 02 Jan 2024
Abstract
Biomedical informatics can be considered as a multidisciplinary research and educational field situated at the intersection of computational sciences (including computer science, data science, mathematics, and statistics), biology, and medicine. In recent years, there have been advances in the field of biomedical informatics.
[...] Read more.
Biomedical informatics can be considered as a multidisciplinary research and educational field situated at the intersection of computational sciences (including computer science, data science, mathematics, and statistics), biology, and medicine. In recent years, there have been advances in the field of biomedical informatics. The current article highlights some interesting state-of-the-art research outcomes in these fields. These include research outcomes in areas like (i) computational biology and medicine, (ii) explainable artificial intelligence (XAI) in biomedical research and clinical practice, (iii) machine learning (including deep learning) methods and application for bioinformatics and healthcare, (iv) imaging informatics, as well as (v) medical statistics and data science. Moreover, the current article also discusses some existing challenges and potential future directions for these research areas to advance the fields of biomedical informatics.
Full article
Open AccessArticle
The Bioinformatics Identification of Potential Protein Glycosylation Genes Associated with a Glioma Stem Cell Signature
by
, , , , , and
BioMedInformatics 2024, 4(1), 75-88; https://doi.org/10.3390/biomedinformatics4010005 - 01 Jan 2024
Abstract
Glioma stem cells (GSCs) contribute to the pathogenesis of glioblastoma (GBM), which is the most malignant form of glioma. The implications and underlying mechanisms of protein glycosylation in GSC phenotypes and GBM malignancy are not fully understood. The implication of protein glycosylation and
[...] Read more.
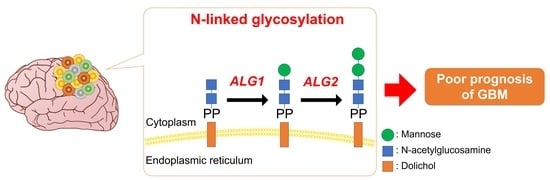
Glioma stem cells (GSCs) contribute to the pathogenesis of glioblastoma (GBM), which is the most malignant form of glioma. The implications and underlying mechanisms of protein glycosylation in GSC phenotypes and GBM malignancy are not fully understood. The implication of protein glycosylation and the corresponding candidate genes on the stem cell properties of GSCs and poor clinical outcomes in GBM were investigated, using datasets from the Gene Expression Omnibus, The Cancer Genome Atlas, and the Chinese Glioma Genome Atlas, accompanied by biological validation in vitro. N-linked glycosylation was significantly associated with GSC properties and the prognosis of GBM in the integrated bioinformatics analyses of clinical specimens. N-linked glycosylation was associated with the glioma grade, molecular biomarkers, and molecular subtypes. The expression levels of the asparagine-linked glycosylation (ALG) enzyme family, which is essential for the early steps in the biosynthesis of N-glycans, were prominently associated with GSC properties and poor survival in patients with GBM with high stem-cell properties. Finally, the oxidative phosphorylation pathway was primarily enriched in GSCs with a high expression of the ALG enzyme family. These findings suggest the role of N-linked glycosylation in the regulation of GSC phenotypes and GBM malignancy.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Survey of Multimodal Medical Question Answering
by
and
BioMedInformatics 2024, 4(1), 50-74; https://doi.org/10.3390/biomedinformatics4010004 - 31 Dec 2023
Abstract
Multimodal medical question answering (MMQA) is a vital area bridging healthcare and Artificial Intelligence (AI). This survey methodically examines the MMQA research published in recent years. We collect academic literature through Google Scholar, applying bibliometric analysis to the publications and datasets used in
[...] Read more.
Multimodal medical question answering (MMQA) is a vital area bridging healthcare and Artificial Intelligence (AI). This survey methodically examines the MMQA research published in recent years. We collect academic literature through Google Scholar, applying bibliometric analysis to the publications and datasets used in these studies. Our analysis uncovers the increasing interest in MMQA over time, with diverse domains such as natural language processing, computer vision, and large language models contributing to the research. The AI methods used in multimodal question answering in the medical domain are a prominent focus, accompanied by applicability of MMQA to the medical field. MMQA in the medical field has its unique challenges due to the sensitive nature of medicine as a science dealing with human health. The survey reveals MMQA research to be in an exploratory stage, discussing different methods, datasets, and potential business models. Future research is expected to focus on application development by big tech companies, such as MedPalm. The survey aims to provide insights into the current state of multimodal medical question answering, highlighting the growing interest from academia and industry. The identified research gaps and trends will guide future investigations and encourage collaborative efforts to advance this transformative field.
Full article
(This article belongs to the Topic Computational Intelligence and Bioinformatics (CIB))
►▼
Show Figures

Figure 1
Open AccessArticle
An Open-Access Dataset of Hospitalized Cardiac-Arrest Patients: Machine-Learning-Based Predictions Using Clinical Documentation
by
, , and
BioMedInformatics 2024, 4(1), 34-49; https://doi.org/10.3390/biomedinformatics4010003 - 27 Dec 2023
Abstract
Cardiac arrest is a sudden loss of heart function with serious consequences. In developing countries, healthcare professionals use clinical documentation to track patient information. These data are used to predict the development of cardiac arrest. We published a dataset through open access to
[...] Read more.
Cardiac arrest is a sudden loss of heart function with serious consequences. In developing countries, healthcare professionals use clinical documentation to track patient information. These data are used to predict the development of cardiac arrest. We published a dataset through open access to advance the research domain. While using this dataset, our work revolved around generating and utilizing synthetic data by harnessing the potential of synthetic data vaults. We conducted a series of experiments by employing state-of-the-art machine-learning techniques. These experiments aimed to assess the performance of our developed predictive model in identifying the likelihood of developing cardiac arrest. This approach was effective in identifying the risk of cardiac arrest in in-patients, even in the absence of electronic medical recording systems. The study evaluated 112 patients who had been transferred from the emergency treatment unit to the cardiac medical ward. The developed model achieved 96% accuracy in predicting the risk of developing cardiac arrest. In conclusion, our study showcased the potential of leveraging clinical documentation and synthetic data to create robust predictive models for cardiac arrest. The outcome of this effort could provide valuable insights and tools for healthcare professionals to preemptively address this critical medical condition.
Full article
(This article belongs to the Section Methods in Biomedical Informatics)
►▼
Show Figures
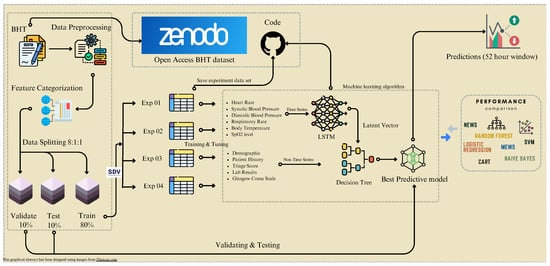
Figure 1
Open AccessArticle
Supporting the Demand on Mental Health Services with AI-Based Conversational Large Language Models (LLMs)
BioMedInformatics 2024, 4(1), 8-33; https://doi.org/10.3390/biomedinformatics4010002 - 22 Dec 2023
Abstract
The demand for psychological counselling has grown significantly in recent years, particularly with the global outbreak of COVID-19, which heightened the need for timely and professional mental health support. Online psychological counselling emerged as the predominant mode of providing services in response to
[...] Read more.
The demand for psychological counselling has grown significantly in recent years, particularly with the global outbreak of COVID-19, which heightened the need for timely and professional mental health support. Online psychological counselling emerged as the predominant mode of providing services in response to this demand. In this study, we propose the Psy-LLM framework, an AI-based assistive tool leveraging large language models (LLMs) for question answering in psychological consultation settings to ease the demand on mental health professions. Our framework combines pre-trained LLMs with real-world professional questions-and-answers (Q&A) from psychologists and extensively crawled psychological articles. The Psy-LLM framework serves as a front-end tool for healthcare professionals, allowing them to provide immediate responses and mindfulness activities to alleviate patient stress. Additionally, it functions as a screening tool to identify urgent cases requiring further assistance. We evaluated the framework using intrinsic metrics, such as perplexity, and extrinsic evaluation metrics, including human participant assessments of response helpfulness, fluency, relevance, and logic. The results demonstrate the effectiveness of the Psy-LLM framework in generating coherent and relevant answers to psychological questions. This article discusses the potential and limitations of using large language models to enhance mental health support through AI technologies.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
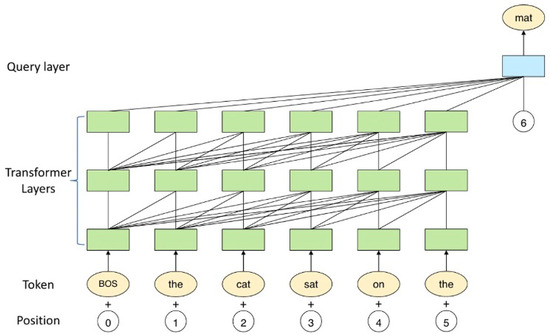
Figure 1
Open AccessEditorial
BioMedInformatics, the Link between Biomedical Informatics, Biology and Computational Medicine
BioMedInformatics 2024, 4(1), 1-7; https://doi.org/10.3390/biomedinformatics4010001 - 21 Dec 2023
Abstract
►▼
Show Figures
Welcome to BioMedInformatics (ISSN: 2673-7426) [...]
Full article

Figure 1
Open AccessArticle
Optimized FIR Filter Using Genetic Algorithms: A Case Study of ECG Signals Filter Optimization
BioMedInformatics 2023, 3(4), 1197-1215; https://doi.org/10.3390/biomedinformatics3040071 - 08 Dec 2023
Abstract
The advancement in technology and the availability of specialized digital signal processing chips have made digital filter design and implementation more feasible in a variety of fields, including biomedical engineering. This paper makes two key contributions. First, it uses a genetic algorithm to
[...] Read more.
The advancement in technology and the availability of specialized digital signal processing chips have made digital filter design and implementation more feasible in a variety of fields, including biomedical engineering. This paper makes two key contributions. First, it uses a genetic algorithm to optimize the coefficients of finite impulse response (FIR) filters. Second, it conducts a case study on using genetic algorithms to optimize FIR filters for electrocardiogram (ECG) biomedical signal noise removal. The goal of the proposed filter design approach is to achieve the desired signal bandwidth while minimizing the side lobe level and eliminating unwanted signals using a genetic algorithm. The results of a comprehensive analysis show that the genetic algorithm-based filter is more effective than conventional filter designs in terms of noise removal efficiency.
Full article
(This article belongs to the Special Issue Application of Semantic Web Technologies in Biomedicine and Biomedical Informatics)
►▼
Show Figures

Figure 1
Open AccessReview
Transforming Drug Design: Innovations in Computer-Aided Discovery for Biosimilar Agents
BioMedInformatics 2023, 3(4), 1178-1196; https://doi.org/10.3390/biomedinformatics3040070 - 08 Dec 2023
Cited by 1
Abstract
In pharmaceutical research and development, pursuing novel therapeutics and optimizing existing drugs have been revolutionized by the fusion of cutting-edge technologies and computational methodologies. Over the past few decades, the field of drug design has undergone a remarkable transformation, catalyzed by the rapid
[...] Read more.
In pharmaceutical research and development, pursuing novel therapeutics and optimizing existing drugs have been revolutionized by the fusion of cutting-edge technologies and computational methodologies. Over the past few decades, the field of drug design has undergone a remarkable transformation, catalyzed by the rapid advancement of computer-aided discovery techniques and the emergence of biosimilar agents. This dynamic interplay between scientific innovation and technological prowess has expedited the drug discovery process and paved the way for more targeted, effective, and personalized treatment approaches. This review investigates the transformative computer-aided discovery techniques for biosimilar agents in reshaping drug design. It examines how computational methods expedite drug candidate identification and explores the rise of cost-effective biosimilars as alternatives to biologics. Through this analysis, this study highlights the potential of these innovations to enhance the efficiency and accessibility of pharmaceutical development. It represents a pioneering effort to examine how computer-aided discovery is revolutionizing biosimilar agent development, exploring its applications, challenges, and prospects.
Full article
(This article belongs to the Special Issue Advances in Structural Bioinformatics and Next-Generation Sequence Analysis for Drug Design)
►▼
Show Figures
Figure 1
Open AccessReview
Genomics for Emerging Pathogen Identification and Monitoring: Prospects and Obstacles
by
, , , , , , , and
BioMedInformatics 2023, 3(4), 1145-1177; https://doi.org/10.3390/biomedinformatics3040069 - 07 Dec 2023
Cited by 1
Abstract
Emerging infectious diseases (EIDs) pose an increasingly significant global burden, driven by urbanization, population explosion, global travel, changes in human behavior, and inadequate public health systems. The recent SARS-CoV-2 pandemic highlights the urgent need for innovative and robust technologies to effectively monitor newly
[...] Read more.
Emerging infectious diseases (EIDs) pose an increasingly significant global burden, driven by urbanization, population explosion, global travel, changes in human behavior, and inadequate public health systems. The recent SARS-CoV-2 pandemic highlights the urgent need for innovative and robust technologies to effectively monitor newly emerging pathogens. Rapid identification, epidemiological surveillance, and transmission mitigation are crucial challenges for ensuring public health safety. Genomics has emerged as a pivotal tool in public health during pandemics, enabling the diagnosis, management, and prediction of infections, as well as the analysis and identification of cross-species interactions and the categorization of infectious agents. Recent advancements in high-throughput DNA sequencing tools have facilitated rapid and precise identification and characterization of emerging pathogens. This review article provides insights into the latest advances in various genomic techniques for pathogen detection and tracking and their applications in global outbreak surveillance. We assess methods that leverage pathogen sequences and explore the role of genomic analysis in understanding the epidemiology of newly emerged infectious diseases. Additionally, we address technical challenges and limitations, ethical and legal considerations, and highlight opportunities for integrating genomics with other surveillance approaches. By delving into the prospects and obstacles of genomics, we can gain valuable insights into its role in mitigating the threats posed by emerging pathogens and improving global preparedness in the face of future outbreaks.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
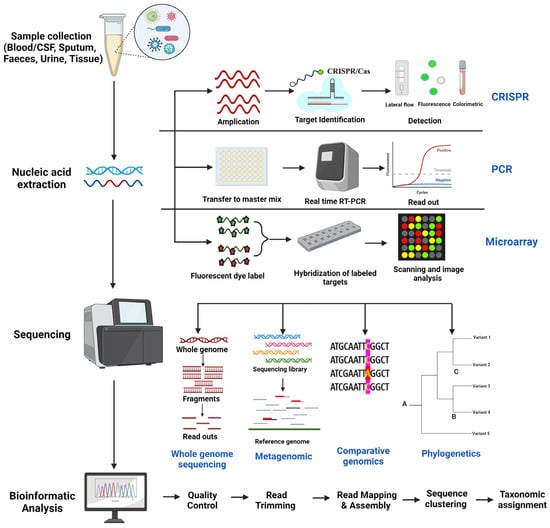
Figure 1
Open AccessArticle
Enhancing Brain Tumor Classification with Transfer Learning across Multiple Classes: An In-Depth Analysis
by
, , , , , , and
BioMedInformatics 2023, 3(4), 1124-1144; https://doi.org/10.3390/biomedinformatics3040068 - 06 Dec 2023
Abstract
This study focuses on leveraging data-driven techniques to diagnose brain tumors through magnetic resonance imaging (MRI) images. Utilizing the rule of deep learning (DL), we introduce and fine-tune two robust frameworks, ResNet 50 and Inception V3, specifically designed for the classification of brain
[...] Read more.
This study focuses on leveraging data-driven techniques to diagnose brain tumors through magnetic resonance imaging (MRI) images. Utilizing the rule of deep learning (DL), we introduce and fine-tune two robust frameworks, ResNet 50 and Inception V3, specifically designed for the classification of brain MRI images. Building upon the previous success of ResNet 50 and Inception V3 in classifying other medical imaging datasets, our investigation encompasses datasets with distinct characteristics, including one with four classes and another with two. The primary contribution of our research lies in the meticulous curation of these paired datasets. We have also integrated essential techniques, including Early Stopping and ReduceLROnPlateau, to refine the model through hyperparameter optimization. This involved adding extra layers, experimenting with various loss functions and learning rates, and incorporating dropout layers and regularization to ensure model convergence in predictions. Furthermore, strategic enhancements, such as customized pooling and regularization layers, have significantly elevated the accuracy of our models, resulting in remarkable classification accuracy. Notably, the pairing of ResNet 50 with the Nadam optimizer yields extraordinary accuracy rates, reaching 99.34% for gliomas, 93.52% for meningiomas, 98.68% for non-tumorous images, and 97.70% for pituitary tumors. These results underscore the transformative potential of our custom-made approach, achieving an aggregate testing accuracy of 97.68% for these four distinct classes. In a two-class dataset, Resnet 50 with the Adam optimizer excels, demonstrating better precision, recall, F1 score, and an overall accuracy of 99.84%. Moreover, it attains perfect per-class accuracy of 99.62% for ‘Tumor Positive’ and 100% for ‘Tumor Negative’, underscoring a remarkable advancement in the realm of brain tumor categorization. This research underscores the innovative possibilities of DL models and our specialized optimization methods in the domain of diagnosing brain cancer from MRI images.
Full article
(This article belongs to the Special Issue Advances in Quantitative Imaging Analysis: From Theory to Practice)
►▼
Show Figures

Figure 1
Open AccessCase Report
Avatar Intervention for Cannabis Use Disorder in a Patient with Schizoaffective Disorder: A Case Report
by
, , , , , , and
BioMedInformatics 2023, 3(4), 1112-1123; https://doi.org/10.3390/biomedinformatics3040067 - 06 Dec 2023
Abstract
Considering the harmful effects of cannabis on individuals with a severe mental disorder and the limited effectiveness of current interventions, this case report showcases the beneficial results of a 10-session Avatar intervention for cannabis use disorder (CUD) on a polysubstance user with a
[...] Read more.
Considering the harmful effects of cannabis on individuals with a severe mental disorder and the limited effectiveness of current interventions, this case report showcases the beneficial results of a 10-session Avatar intervention for cannabis use disorder (CUD) on a polysubstance user with a comorbid schizoaffective disorder. Virtual reality allowed the creation of an Avatar representing a person significantly related to the patient’s drug use. Avatar intervention for CUD aims to combine exposure, relational, and cognitive behavioral therapies while practicing real-life situations and learning how to manage negative emotions and cravings. Throughout therapy and later on, Mr. C managed to maintain abstinence from all substances. Also, an improvement in the severity of CUD, as well as a greater motivation to change consumption, was observed after therapy. As observed by his mother, his psychiatrist, and himself, the benefits of Avatar intervention for CUD extended to other spheres of his life. The drastic results observed in this patient could be promising as an alternative to the current treatment available for people with a dual diagnosis of cannabis use disorder and psychotic disorder, which generally lack effectiveness. A single-blind randomized control trial comparing the treatment with a classical intervention in a larger sample is currently underway to evaluate whether the results are reproducible on a larger sample.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
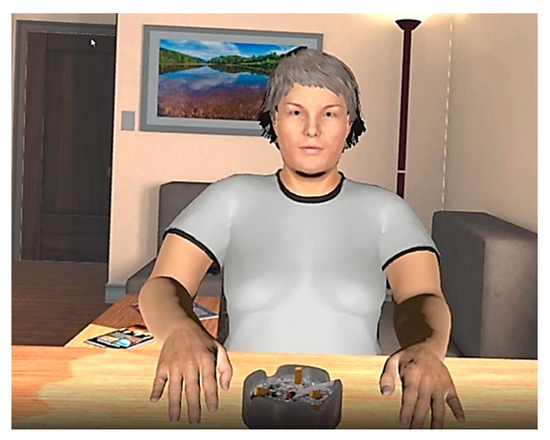

Figure 1
Open AccessReview
Deciphering the Mosaic of Therapeutic Potential: A Scoping Review of Neural Network Applications in Psychotherapy Enhancements
BioMedInformatics 2023, 3(4), 1101-1111; https://doi.org/10.3390/biomedinformatics3040066 - 01 Dec 2023
Abstract
Background: Psychotherapy is a component of the therapeutic options accessible in mental health. Along with psychotherapy techniques and indications, there is a body of studies on what are known as psychotherapy’s common factors. However, up to 40% of patients do not respond to
[...] Read more.
Background: Psychotherapy is a component of the therapeutic options accessible in mental health. Along with psychotherapy techniques and indications, there is a body of studies on what are known as psychotherapy’s common factors. However, up to 40% of patients do not respond to therapy. Artificial intelligence approaches are hoped to enhance this and with the growing body of evidence of the use of neural networks (NNs) in other areas of medicine, this domain is lacking in the field of psychotherapy. This study aims to identify the different uses of NNs in the field of psychotherapy. Methods: A scoping review was conducted in the electronic databases EMBASE, MEDLINE, APA, and CINAHL. The Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement influenced this study’s design. Studies were included if they applied a neural network algorithm in the context of a psychotherapeutic approach. Results: A total of 157 studies were screened for eligibility, of which 32 were fully assessed. Finally, eight articles were analyzed, and three uses were identified: predicting the therapeutic outcomes, content analysis, and automated categorization of psychotherapeutic interactions. Conclusions: Uses of NNs were identified with limited evidence of their effects. The potential implications of these uses could assist the therapist in providing a more personalized therapeutic approach to their patients. Given the paucity of literature, this study provides a path for future research to better understand the efficacy of such uses.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
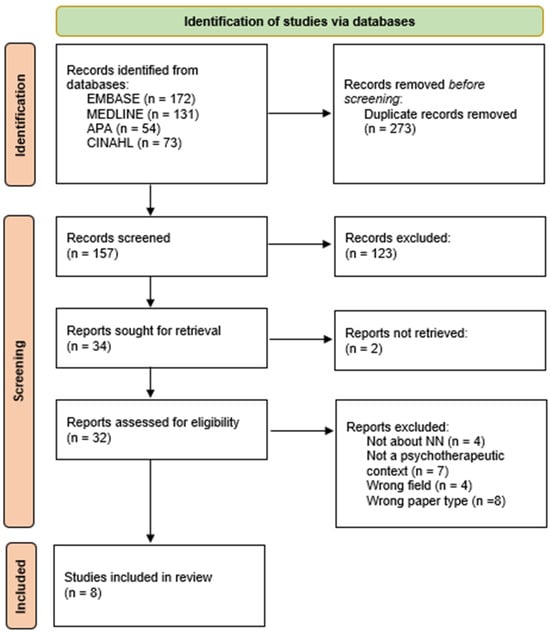
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomedicines, BioMedInformatics, Cancers, JCM, Cells, Current Oncology
Cancer Immunity and Immunotherapy: Early Detection, Diagnosis, Systemic Treatments, Novel Biomarkers, and Resistance Mechanisms
Topic Editors: Tao Jiang, Yongchang ZhangDeadline: 20 April 2024
Topic in
BioMedInformatics, Cancers, Current Oncology, Hemato, Hematology Reports
Leukemia-Challenges and Current Treatment Options
Topic Editors: Giovanni Martinelli, Claudio CerchioneDeadline: 20 June 2024
Topic in
Algorithms, BDCC, BioMedInformatics, Information, Mathematics
Machine Learning Empowered Drug Screen
Topic Editors: Teng Zhou, Jiaqi Wang, Youyi SongDeadline: 31 August 2024
Topic in
Biology, BioMedInformatics, Cancers, Genes, IJMS
The 22nd International Conference on Bioinformatics (InCoB 2023): Translational Bioinformatics Transforming Life
Topic Editors: Jyotsna Batra, Srilakshmi Srinivasan, Shoba Ranganathan, Asif M. Khan, Harpreet SinghDeadline: 15 November 2024

Conferences
Special Issues
Special Issue in
BioMedInformatics
Feature Papers in Medical Statistics and Data Science Section
Guest Editor: Pentti NieminenDeadline: 30 June 2024
Special Issue in
BioMedInformatics
Feature Papers in Computational Biology and Medicine
Guest Editor: Hans BinderDeadline: 31 July 2024
Special Issue in
BioMedInformatics
Advances in Quantitative Imaging Analysis: From Theory to Practice
Guest Editors: Federico Mastroleo, Angela Ammirabile, Giulia MarvasoDeadline: 30 November 2024
Special Issue in
BioMedInformatics
Deep Learning Methods and Application for Bioinformatics and Healthcare
Guest Editors: Pan Zheng, Bin WangDeadline: 31 December 2024













.jpg)



