Journal Description
Epigenomes
Epigenomes
is an international, peer-reviewed, open access journal on epigenetics and epigenomics, published quarterly online by MDPI. The Epigenetics Society is affiliated with Epigenomes and its members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PMC, PubMed, Embase, PubAg, CAPlus / SciFinder, and other databases.
- Journal Rank: CiteScore - Q2 (Biochemistry, Genetics and Molecular Biology (miscellaneous))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.3 days after submission; acceptance to publication is undertaken in 3.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
2.5 (2022);
5-Year Impact Factor:
2.1 (2022)
Latest Articles
The Genomic Shock Hypothesis: Genetic and Epigenetic Alterations of Transposable Elements after Interspecific Hybridization in Plants
Epigenomes 2024, 8(1), 2; https://doi.org/10.3390/epigenomes8010002 - 27 Dec 2023
Abstract
Transposable elements (TEs) are major components of plant genomes with the ability to change their position in the genome or to create new copies of themselves in other positions in the genome. These can cause gene disruption and large-scale genomic alterations, including inversions,
[...] Read more.
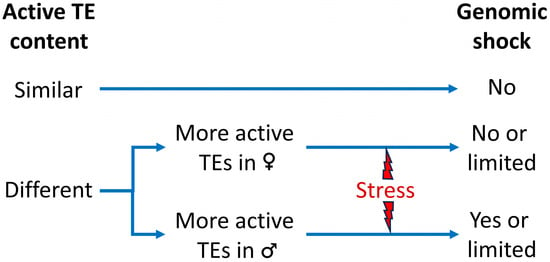
Transposable elements (TEs) are major components of plant genomes with the ability to change their position in the genome or to create new copies of themselves in other positions in the genome. These can cause gene disruption and large-scale genomic alterations, including inversions, deletions, and duplications. Host organisms have evolved a set of mechanisms to suppress TE activity and counter the threat that they pose to genome integrity. These includes the epigenetic silencing of TEs mediated by a process of RNA-directed DNA methylation (RdDM). In most cases, the silencing machinery is very efficient for the vast majority of TEs. However, there are specific circumstances in which TEs can evade such silencing mechanisms, for example, a variety of biotic and abiotic stresses or in vitro culture. Hybridization is also proposed as an inductor of TE proliferation. In fact, the discoverer of the transposons, Barbara McClintock, first hypothesized that interspecific hybridization provides a “genomic shock” that inhibits the TE control mechanisms leading to the mobilization of TEs. However, the studies carried out on this topic have yielded diverse results, showing in some cases a total absence of mobilization or being limited to only some TE families. Here, we review the current knowledge about the impact of interspecific hybridization on TEs in plants and the possible implications of changes in the epigenetic mechanisms.
Full article
(This article belongs to the Collection Epigenetic Control in Plants)
►
Show Figures
Open AccessReview
Inheritance of Stress Responses via Small Non-Coding RNAs in Invertebrates and Mammals
by
and
Epigenomes 2024, 8(1), 1; https://doi.org/10.3390/epigenomes8010001 - 19 Dec 2023
Abstract
While reports on the generational inheritance of a parental response to stress have been widely reported in animals, the molecular mechanisms behind this phenomenon have only recently emerged. The booming interest in epigenetic inheritance has been facilitated in part by the discovery that
[...] Read more.
While reports on the generational inheritance of a parental response to stress have been widely reported in animals, the molecular mechanisms behind this phenomenon have only recently emerged. The booming interest in epigenetic inheritance has been facilitated in part by the discovery that small non-coding RNAs are one of its principal conduits. Discovered 30 years ago in the Caenorhabditis elegans nematode, these small molecules have since cemented their critical roles in regulating virtually all aspects of eukaryotic development. Here, we provide an overview on the current understanding of epigenetic inheritance in animals, including mice and C. elegans, as it pertains to stresses such as temperature, nutritional, and pathogenic encounters. We focus on C. elegans to address the mechanistic complexity of how small RNAs target their cohort mRNAs to effect gene expression and how they govern the propagation or termination of generational perdurance in epigenetic inheritance. Presently, while a great amount has been learned regarding the heritability of gene expression states, many more questions remain unanswered and warrant further investigation.
Full article
(This article belongs to the Special Issue Transgenerational Epigenetic Inheritance)
►▼
Show Figures
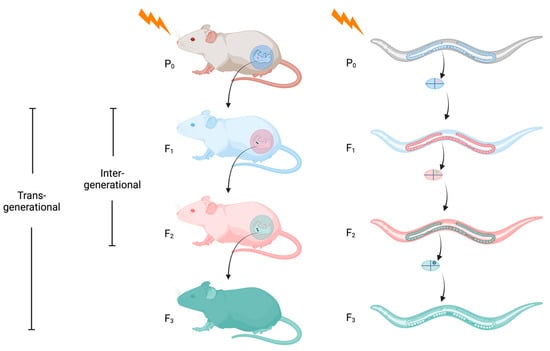
Figure 1
Open AccessReview
Epigenetic Mechanisms in Hematologic Aging and Premalignant Conditions
Epigenomes 2023, 7(4), 32; https://doi.org/10.3390/epigenomes7040032 - 12 Dec 2023
Cited by 1
Abstract
Hematopoietic stem cells (HSCs) are essential for maintaining overall health by continuously generating blood cells throughout an individual’s lifespan. However, as individuals age, the hematopoietic system undergoes significant functional decline, rendering them more susceptible to age-related diseases. Growing research evidence has highlighted the
[...] Read more.
Hematopoietic stem cells (HSCs) are essential for maintaining overall health by continuously generating blood cells throughout an individual’s lifespan. However, as individuals age, the hematopoietic system undergoes significant functional decline, rendering them more susceptible to age-related diseases. Growing research evidence has highlighted the critical role of epigenetic regulation in this age-associated decline. This review aims to provide an overview of the diverse epigenetic mechanisms involved in the regulation of normal HSCs during the aging process and their implications in aging-related diseases. Understanding the intricate interplay of epigenetic mechanisms that contribute to aging-related changes in the hematopoietic system holds great potential for the development of innovative strategies to delay the aging process. In fact, interventions targeting epigenetic modifications have shown promising outcomes in alleviating aging-related phenotypes and extending lifespan in various animal models. Small molecule-based therapies and reprogramming strategies enabling epigenetic rejuvenation have emerged as effective approaches for ameliorating or even reversing aging-related conditions. By acquiring a deeper understanding of these epigenetic mechanisms, it is anticipated that interventions can be devised to prevent or mitigate the rates of hematologic aging and associated diseases later in life. Ultimately, these advancements have the potential to improve overall health and enhance the quality of life in aging individuals.
Full article
(This article belongs to the Special Issue Epigenetics in Hematologic Malignancies)
►▼
Show Figures
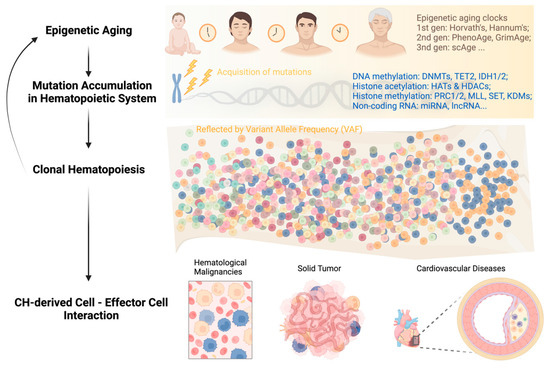
Figure 1
Open AccessReview
World Trade Center Exposure, DNA Methylation Changes, and Cancer: A Review of Current Evidence
by
, , , , , , , , , , and
Epigenomes 2023, 7(4), 31; https://doi.org/10.3390/epigenomes7040031 - 08 Dec 2023
Abstract
Introduction: Known carcinogens in the dust and fumes from the destruction of the World Trade Center (WTC) towers on 9 November 2001 included metals, asbestos, and organic pollutants, which have been shown to modify epigenetic status. Epigenome-wide association analyses (EWAS) using uniform
[...] Read more.
Introduction: Known carcinogens in the dust and fumes from the destruction of the World Trade Center (WTC) towers on 9 November 2001 included metals, asbestos, and organic pollutants, which have been shown to modify epigenetic status. Epigenome-wide association analyses (EWAS) using uniform (Illumina) methodology have identified novel epigenetic profiles of WTC exposure. Methods: We reviewed all published data, comparing differentially methylated gene profiles identified in the prior EWAS studies of WTC exposure. This included DNA methylation changes in blood-derived DNA from cases of cancer-free “Survivors” and those with breast cancer, as well as tissue-derived DNA from “Responders” with prostate cancer. Emerging molecular pathways related to the observed DNA methylation changes in WTC-exposed groups were explored and summarized. Results: WTC dust exposure appears to be associated with DNA methylation changes across the genome. Notably, WTC dust exposure appears to be associated with increased global DNA methylation; direct dysregulation of cancer genes and pathways, including inflammation and immune system dysregulation; and endocrine system disruption, as well as disruption of cholesterol homeostasis and lipid metabolism. Conclusion: WTC dust exposure appears to be associated with biologically meaningful DNA methylation changes, with implications for carcinogenesis and development of other chronic diseases.
Full article
(This article belongs to the Special Issue Environmental Epigenomes)
Open AccessArticle
Comparison of Yersinia enterocolitica DNA Methylation at Ambient and Host Temperatures
Epigenomes 2023, 7(4), 30; https://doi.org/10.3390/epigenomes7040030 - 30 Nov 2023
Abstract
Pathogenic bacteria recognize environmental cues to vary gene expression for host adaptation. Moving from ambient to host temperature, Yersinia enterocolitica responds by immediately repressing flagella synthesis and inducing the virulence plasmid (pYV)-encoded type III secretion system. In contrast, shifting from host to ambient temperature
[...] Read more.
Pathogenic bacteria recognize environmental cues to vary gene expression for host adaptation. Moving from ambient to host temperature, Yersinia enterocolitica responds by immediately repressing flagella synthesis and inducing the virulence plasmid (pYV)-encoded type III secretion system. In contrast, shifting from host to ambient temperature requires 2.5 generations to restore motility, suggesting a link to the cell cycle. We hypothesized that differential DNA methylation contributes to temperature-regulated gene expression. We tested this hypothesis by comparing single-molecule real-time (SMRT) sequencing of Y. enterocolitica DNA from cells growing exponentially at 22 °C and 37 °C. The inter-pulse duration ratio rather than the traditional QV scoring was the kinetic metric to compare DNA from cells grown at each temperature. All 565 YenI restriction sites were fully methylated at both temperatures. Among the 27,118 DNA adenine methylase (Dam) sites, 42 had differential methylation patterns, while 17 remained unmethylated regardless of the temperature. A subset of the differentially methylated Dam sites localized to promoter regions of predicted regulatory genes including LysR-type and PadR-like transcriptional regulators and a cyclic-di-GMP phosphodiesterase. The unmethylated Dam sites localized with a bias to the replication terminus, suggesting they were protected from Dam methylase. No cytosine methylation was detected at Dcm sites.
Full article
(This article belongs to the Special Issue Epigenetic and Epitranscriptomic Determinants of Host-Microbe Interactions)
►▼
Show Figures
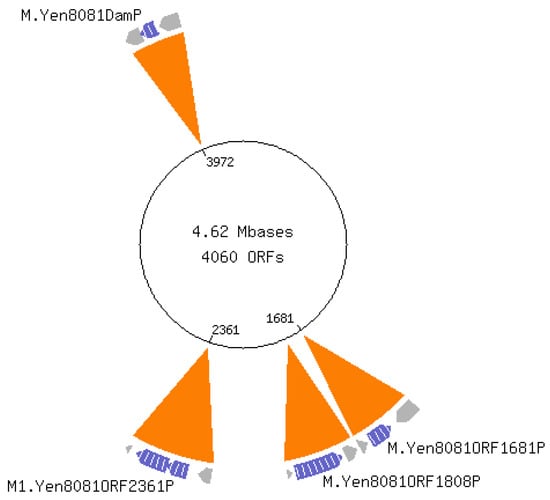
Figure 1
Open AccessReview
Out of the Silence: Insights into How Genes Escape X-Chromosome Inactivation
Epigenomes 2023, 7(4), 29; https://doi.org/10.3390/epigenomes7040029 - 23 Nov 2023
Abstract
The silencing of all but one X chromosome in mammalian cells is a remarkable epigenetic process leading to near dosage equivalence in X-linked gene products between the sexes. However, equally remarkable is the ability of a subset of genes to continue to be
[...] Read more.
The silencing of all but one X chromosome in mammalian cells is a remarkable epigenetic process leading to near dosage equivalence in X-linked gene products between the sexes. However, equally remarkable is the ability of a subset of genes to continue to be expressed from the otherwise inactive X chromosome—in some cases constitutively, while other genes are variable between individuals, tissues or cells. In this review we discuss the advantages and disadvantages of the approaches that have been used to identify escapees. The identity of escapees provides important clues to mechanisms underlying escape from XCI, an arena of study now moving from correlation to functional studies. As most escapees show greater expression in females, the not-so-inactive X chromosome is a substantial contributor to sex differences in humans, and we highlight some examples of such impact.
Full article
(This article belongs to the Special Issue X-Chromosome Inactivation)
►▼
Show Figures

Graphical abstract
Open AccessReview
IndiSPENsable for X Chromosome Inactivation and Gene Silencing
by
and
Epigenomes 2023, 7(4), 28; https://doi.org/10.3390/epigenomes7040028 - 02 Nov 2023
Abstract
For about 30 years, SPEN has been the subject of research in many different fields due to its variety of functions and its conservation throughout a wide spectrum of species, like worms, arthropods, and vertebrates. To date, 216 orthologues have been documented. SPEN
[...] Read more.
For about 30 years, SPEN has been the subject of research in many different fields due to its variety of functions and its conservation throughout a wide spectrum of species, like worms, arthropods, and vertebrates. To date, 216 orthologues have been documented. SPEN had been studied for its role in gene regulation in the context of cell signaling, including the NOTCH or nuclear hormone receptor signaling pathways. More recently, SPEN has been identified as a major regulator of initiation of chromosome-wide gene silencing during X chromosome inactivation (XCI) in mammals, where its function remains to be fully understood. Dependent on the biological context, SPEN functions via mechanisms which include different domains. While some domains of SPEN are highly conserved in sequence and secondary structure, species-to-species differences exist that might lead to mechanistic differences. Initiation of XCI appears to be different between humans and mice, which raises additional questions about the extent of generalization of SPEN’s function in XCI. In this review, we dissect the mechanism of SPEN in XCI. We discuss its subregions and domains, focusing on its role as a major regulator. We further highlight species-related research, specifically of mouse and human SPEN, with the aim to reveal and clarify potential species-to-species differences in SPEN’s function.
Full article
(This article belongs to the Special Issue X-Chromosome Inactivation)
►▼
Show Figures

Figure 1
Open AccessArticle
Stress and DNA Methylation of Blood Leukocytes among Pregnant Latina Women
by
, , , , , , , and
Epigenomes 2023, 7(4), 27; https://doi.org/10.3390/epigenomes7040027 - 01 Nov 2023
Abstract
Latinas experience physical and psychological stressors in pregnancy leading to increased morbidity and higher risk for adverse birth outcomes. Epigenetic changes, including DNA methylation (DNAm), have been proposed as markers to create more refined risk stratification, yet few of these studies have examined
[...] Read more.
Latinas experience physical and psychological stressors in pregnancy leading to increased morbidity and higher risk for adverse birth outcomes. Epigenetic changes, including DNA methylation (DNAm), have been proposed as markers to create more refined risk stratification, yet few of these studies have examined these changes in Latinas. We conducted a secondary analysis of stored blood leukocytes of Latina women (n = 58) enrolled in a larger National Institutes of Health funded R01 project (2011–2016). We examined DNAm on eight candidate stress genes to compare physically and psychologically stressed participants to healthy (low stress) participants. We found unique CpGs that were differentially methylated in stressed women early- and mid-pregnancy compared to the healthy group, though none remained significant after FDR correction. Both physical and psychological stress were associated with hypomethylation at two consecutive CpG sites on NR3C1 in early pregnancy and one CpG site on NR3C1 in mid-pregnancy before adjustment. Stress was also associated with hypomethylation at two CpG sites on FKBP5 in early and mid-pregnancy but were no longer significant after FDR adjustment. Though we did not find statistically significant differences in DNAm during pregnancy between stressed and healthy women in this sample, signals were consistent with previous findings. Future work in larger samples should further examine the associations between stress and DNAm in pregnancy as this mechanism may explain underlying perinatal health inequities.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures
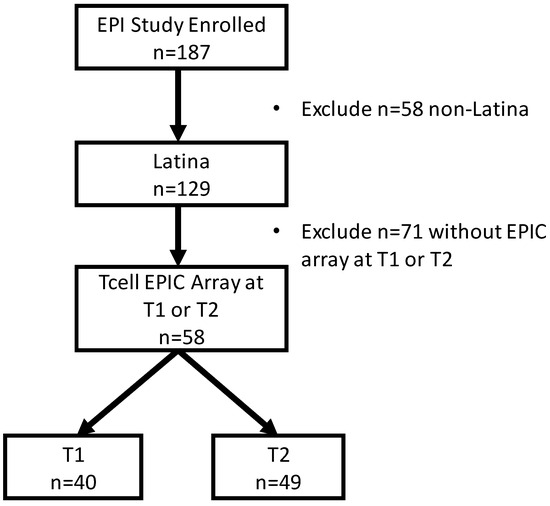
Figure 1
Open AccessCorrection
Correction: Islam, R.A.; Rallis, C. Ribosomal Biogenesis and Heterogeneity in Development, Disease, and Aging. Epigenomes 2023, 7, 17
Epigenomes 2023, 7(4), 26; https://doi.org/10.3390/epigenomes7040026 - 26 Oct 2023
Abstract
23. Akirtava, C.; May, G.E.; McManus, C.J. False-Positive IRESes from Hoxa9 and
Other Genes Resulting from Errors in Mam-malian 5’ UTR Annotations [...] Full article
Other Genes Resulting from Errors in Mam-malian 5’ UTR Annotations [...] Full article
Open AccessArticle
Maternal MicroRNA Profile Changes When LH Is Added to the Ovarian Stimulation Protocol: A Pilot Study
by
, , , , , and
Epigenomes 2023, 7(4), 25; https://doi.org/10.3390/epigenomes7040025 - 06 Oct 2023
Abstract
While the use of follicle-stimulating hormone (FSH) in ovarian stimulation for in vitro fertilization (IVF) is an established practice, the use of luteinizing hormone (LH) remains debatable. MicroRNAs (miRNAs) are short, endogenous, non-coding transcripts that control a variety of cellular functions, such as
[...] Read more.
While the use of follicle-stimulating hormone (FSH) in ovarian stimulation for in vitro fertilization (IVF) is an established practice, the use of luteinizing hormone (LH) remains debatable. MicroRNAs (miRNAs) are short, endogenous, non-coding transcripts that control a variety of cellular functions, such as gonadotrophin production and follicular development. The goal of this pilot study was to investigate whether the employment of recombinant LH (rLH) in ovarian stimulation protocols results in changes in the miRNA profiles in human oocytes. Patients were divided into two groups: seven received recombinant FSH (rFSH, 225 IU), and six received rFSH (150 IU) plus rLH (75 IU). MiRNA predesigned panels and real-time PCR technology were used to analyze the oocytes retrieved from the follicular ovarian retrieval. Among the miRNAs evaluated, a series of them evidenced upregulation or downregulation in their expression in the FSH plus LH group compared to the FSH group. Considering the results obtained from the functional and network analysis, the different maternal miRNA profiles in the two groups revealed a differential modulation of pathways involved in numerous biological functions. Overall, based on the pathways associated with most of these maternal miRNAs, the presence of LH may result in a different modulation of pathways regulating survival under the control of a Tp53-related mechanism. Interestingly, among the miRNAs differentially expressed in oocytes of the two groups, we have found miRNAs already investigated at ovarian, follicular, oocyte, and embryonic levels: hsa-miR-484, hsa-miR-222, hsa-miR-520d-5p, hsa-miRNA-17, hsa-miR-548, and hsa-miR-140. Thus, investigation into the role of these miRNAs in oocyte molecular pathways may help determine how LH affects oocyte competence and eventually leads to the clinical improvement of IVF.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessReview
The ErbB Signaling Network and Its Potential Role in Endometrial Cancer
by
, , , , , and
Epigenomes 2023, 7(4), 24; https://doi.org/10.3390/epigenomes7040024 - 01 Oct 2023
Cited by 2
Abstract
Endometrial cancer (EC) is the second most common malignancy of the female reproductive system worldwide. The updated EC classification emphasizes the significant role of various signaling pathways such as PIK3CA-PIK3R1-PTEN and RTK/RAS/β-catenin in EC pathogenesis. Some of these pathways are part of the
[...] Read more.
Endometrial cancer (EC) is the second most common malignancy of the female reproductive system worldwide. The updated EC classification emphasizes the significant role of various signaling pathways such as PIK3CA-PIK3R1-PTEN and RTK/RAS/β-catenin in EC pathogenesis. Some of these pathways are part of the EGF system signaling network, which becomes hyperactivated by various mechanisms and participates in cancer pathogenesis. In EC, the expression of ErbB receptors is significantly different, compared with the premenopausal and postmenopausal endometrium, mainly because of the increased transcriptional activity of ErbB encoding genes in EC cells. Moreover, there are some differences in ErbB-2 receptor profile among EC subgroups that could be explained by the alterations in pathophysiology and clinical behavior of various EC histologic subtypes. The fact that ErbB-2 receptor expression is more common in aggressive EC histologic subtypes (papillary serous and clear cell) could indicate a future role of ErbB-targeted therapies in well-defined EC subgroups with overexpression of ErbB receptors.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures

Figure 1
Open AccessReview
Current Approaches to Epigenetic Therapy
by
, , , , , , , , , and
Epigenomes 2023, 7(4), 23; https://doi.org/10.3390/epigenomes7040023 - 30 Sep 2023
Abstract
►▼
Show Figures
Epigenetic therapy is a promising tool for the treatment of a wide range of diseases. Several fundamental epigenetic approaches have been proposed. Firstly, the use of small molecules as epigenetic effectors, as the most developed pharmacological method, has contributed to the introduction of
[...] Read more.
Epigenetic therapy is a promising tool for the treatment of a wide range of diseases. Several fundamental epigenetic approaches have been proposed. Firstly, the use of small molecules as epigenetic effectors, as the most developed pharmacological method, has contributed to the introduction of a number of drugs into clinical practice. Secondly, various innovative epigenetic approaches based on dCas9 and the use of small non-coding RNAs as therapeutic agents are also under extensive research. In this review, we present the current state of research in the field of epigenetic therapy, considering the prospects for its application and possible limitations.
Full article

Figure 1
Open AccessArticle
Linc2function: A Comprehensive Pipeline and Webserver for Long Non-Coding RNA (lncRNA) Identification and Functional Predictions Using Deep Learning Approaches
by
, , , , and
Epigenomes 2023, 7(3), 22; https://doi.org/10.3390/epigenomes7030022 - 15 Sep 2023
Abstract
Long non-coding RNAs (lncRNAs), comprising a significant portion of the human transcriptome, serve as vital regulators of cellular processes and potential disease biomarkers. However, the function of most lncRNAs remains unknown, and furthermore, existing approaches have focused on gene-level investigation. Our work emphasizes
[...] Read more.
Long non-coding RNAs (lncRNAs), comprising a significant portion of the human transcriptome, serve as vital regulators of cellular processes and potential disease biomarkers. However, the function of most lncRNAs remains unknown, and furthermore, existing approaches have focused on gene-level investigation. Our work emphasizes the importance of transcript-level annotation to uncover the roles of specific transcript isoforms. We propose that understanding the mechanisms of lncRNA in pathological processes requires solving their structural motifs and interactomes. A complete lncRNA annotation first involves discriminating them from their coding counterparts and then predicting their functional motifs and target bio-molecules. Current in silico methods mainly perform primary-sequence-based discrimination using a reference model, limiting their comprehensiveness and generalizability. We demonstrate that integrating secondary structure and interactome information, in addition to using transcript sequence, enables a comprehensive functional annotation. Annotating lncRNA for newly sequenced species is challenging due to inconsistencies in functional annotations, specialized computational techniques, limited accessibility to source code, and the shortcomings of reference-based methods for cross-species predictions. To address these challenges, we developed a pipeline for identifying and annotating transcript sequences at the isoform level. We demonstrate the effectiveness of the pipeline by comprehensively annotating the lncRNA associated with two specific disease groups. The source code of our pipeline is available under the MIT licensefor local use by researchers to make new predictions using the pre-trained models or to re-train models on new sequence datasets. Non-technical users can access the pipeline through a web server setup.
Full article
(This article belongs to the Collection Feature Papers in Epigenomes)
►▼
Show Figures
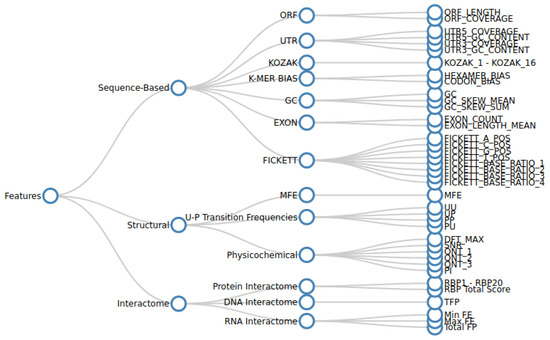
Figure 1
Open AccessEditorial
Environmental Epigenomes
Epigenomes 2023, 7(3), 21; https://doi.org/10.3390/epigenomes7030021 - 07 Sep 2023
Abstract
Research in epigenetics has dramatically risen during the last decade to include aspects of environmental biology. However, many questions remain regarding the effects of environmental stressors on the epigenome, incorporating the particular role of epigenetic mechanisms in the adaptation and evolution of organisms
[...] Read more.
Research in epigenetics has dramatically risen during the last decade to include aspects of environmental biology. However, many questions remain regarding the effects of environmental stressors on the epigenome, incorporating the particular role of epigenetic mechanisms in the adaptation and evolution of organisms in changing environments. Epigenetics is commonly defined as mitotically and/or meiotically heritable changes in gene function that occur without altering the underlying DNA sequence. It encompasses DNA (hydroxy)methylation, histone modifications, chromatin structure, and non-coding RNAs that may be inherited across generations under certain circumstances. Epigenetic mechanisms are perfect candidates to extend our understanding of the impact of environmental stressors on organisms and to explain the rapid phenomenon of adaptive evolution. Existing evidence shows that environmental cues can affect the epigenome and modify gene expression accordingly. These changes can then induce phenotypic modifications that are morphological, physiological, or behavioral at the organismal level. In this Special Issue focusing on environmental epigenetics, we provide an overview of influences to the epigenome that are driven by various environmental and evolutionary factors, with a particular focus on DNA methylation (DNAm). Five research groups have contributed insightful studies or reviews on (1) DNAm and demethylation events affected by the exposome; (2) DNAm as a potential biomarker to determine cardiometabolic risk early in life; (3) consequences of DNAm across multiple generations; (4) DNAm variation within natural animal populations; and (5) epigenetic mechanisms in genetically uniform organisms. Collectively, the articles from this Special Issue consistently support that environmental changes can induce long-lasting epigenetic effects within a given organism pertaining to individual risk for disease, or multi-generational impacts that ultimately impact evolution.
Full article
(This article belongs to the Special Issue Environmental Epigenomes)
Open AccessFeature PaperReview
Integrated Multimodal Omics and Dietary Approaches for the Management of Neurodegeneration
by
and
Epigenomes 2023, 7(3), 20; https://doi.org/10.3390/epigenomes7030020 - 01 Sep 2023
Abstract
Neurodegenerative diseases, such as Alzheimer’s disease and Parkinson’s disease, are caused by a combination of multiple events that damage neuronal function. A well-characterized biomarker of neurodegeneration is the accumulation of proteinaceous aggregates in the brain. However, the gradually worsening symptoms of neurodegenerative diseases
[...] Read more.
Neurodegenerative diseases, such as Alzheimer’s disease and Parkinson’s disease, are caused by a combination of multiple events that damage neuronal function. A well-characterized biomarker of neurodegeneration is the accumulation of proteinaceous aggregates in the brain. However, the gradually worsening symptoms of neurodegenerative diseases are unlikely to be solely due to the result of a mutation in a single gene, but rather a multi-step process involving epigenetic changes. Recently, it has been suggested that a fraction of epigenetic alternations may be correlated to neurodegeneration in the brain. Unlike DNA mutations, epigenetic alterations are reversible, and therefore raise the possibilities for therapeutic intervention, including dietary modifications. Additionally, reactive oxygen species may contribute to the pathogenesis of Alzheimer’s disease and Parkinson’s disease through epigenetic alternation. Given that the antioxidant properties of plant-derived phytochemicals are likely to exhibit pleiotropic effects against ROS-mediated epigenetic alternation, dietary intervention may be promising for the management of neurodegeneration in these diseases. In this review, the state-of-the-art applications using single-cell multimodal omics approaches, including epigenetics, and dietary approaches for the identification of novel biomarkers and therapeutic approaches for the treatment of neurodegenerative diseases are discussed.
Full article
(This article belongs to the Special Issue Epigenetics of Neurobiology and Brain Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Multiomics Data Analysis Identified CpG Sites That Mediate the Impact of Smoking on Cardiometabolic Traits
by
Epigenomes 2023, 7(3), 19; https://doi.org/10.3390/epigenomes7030019 - 22 Aug 2023
Abstract
►▼
Show Figures
Understanding the epigenome paths through which smoking contributes to cardiometabolic traits is important for downstream applications. In this study, an SNP-based analytical pipeline was used to integrate several publicly available datasets in order to identify CpG sites that mediate the impact of smoking
[...] Read more.
Understanding the epigenome paths through which smoking contributes to cardiometabolic traits is important for downstream applications. In this study, an SNP-based analytical pipeline was used to integrate several publicly available datasets in order to identify CpG sites that mediate the impact of smoking on cardiometabolic traits and to investigate the underlying molecular mechanisms. After applying stringent statistical criteria, 11 CpG sites were detected that showed significant association (p < 5 × 10−8) with cardiometabolic traits at both the discovery and replication stages. By integrating eQTL data, I found genes behind a number of these associations. cg05228408 was hypomethylated in smokers and contributed to higher blood pressure by lowering the expression of the CLCN6 gene. cg08639339 was hypermethylated in smokers and lowered the metabolic rate by increasing the expression of RAB29; furthermore, I noted TMEM120A mediated the impact of smoking-cg17325771 on LDL, and LTBP3 mediated the smoking-cg07029024 effect on heart rate. The pathway analysis identified processes through which the identified genes impact their traits. This study provides a list of CpG sites that mediates the impact of smoking on cardiometabolic traits and a framework to investigate the underlying molecular paths using publicly available data.
Full article

Figure 1
Open AccessArticle
Excessive Gestational Weight Gain Alters DNA Methylation and Influences Foetal and Neonatal Body Composition
by
, , , , and
João Victor da Silva Guerra
Epigenomes 2023, 7(3), 18; https://doi.org/10.3390/epigenomes7030018 - 16 Aug 2023
Abstract
►▼
Show Figures
Background: Changes in body weight are associated with the regulation of DNA methylation (DNAm). In this study, we investigated the associations between maternal gestational weight gain-related DNAm and foetal and neonatal body composition. Methods: Brazilian pregnant women from the Araraquara Cohort Study were
[...] Read more.
Background: Changes in body weight are associated with the regulation of DNA methylation (DNAm). In this study, we investigated the associations between maternal gestational weight gain-related DNAm and foetal and neonatal body composition. Methods: Brazilian pregnant women from the Araraquara Cohort Study were followed up during pregnancy, delivery, and after hospital discharge. Women with normal pre-pregnancy BMI were allocated into two groups: adequate gestational weight gain (AGWG, n = 45) and excessive gestational weight gain (EGWG, n = 30). Foetal and neonatal body composition was evaluated via ultrasound and plethysmography, respectively. DNAm was assessed in maternal blood using Illumina Infinium MethylationEPIC BeadChip arrays. Linear regression models were used to explore the associations between DNAm and foetal and neonatal body composition. Results: Maternal weight, GWG, neonatal weight, and fat mass were higher in the EGWG group. Analysis of DNAm identified 46 differentially methylated positions and 11 differentially methylated regions (DMRs) between the EGWG and AGWG groups. Nine human phenotypes were enriched for these 11 DMRs located in 13 genes (EMILIN1, HOXA5, CPT1B, CLDN9, ZFP57, BRCA1, POU5F1, ANKRD33, HLA-B, RANBP17, ZMYND11, DIP2C, TMEM232), highlighting the terms insulin resistance, and hyperglycaemia. Maternal DNAm was associated with foetal total thigh and arm tissues and subcutaneous thigh and arm fat, as well as with neonatal fat mass percentage and fat mass. Conclusion: The methylation pattern in the EGWG group indicated a risk for developing chronic diseases and involvement of maternal DNAm in foetal lean and fat mass and in neonatal fat mass.
Full article
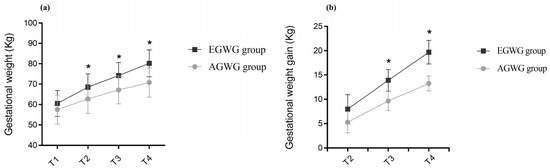
Figure 1
Open AccessReview
Ribosomal Biogenesis and Heterogeneity in Development, Disease, and Aging
Epigenomes 2023, 7(3), 17; https://doi.org/10.3390/epigenomes7030017 - 11 Aug 2023
Cited by 1
Abstract
►▼
Show Figures
Although reported in the literature, ribosome heterogeneity is a phenomenon whose extent and implications in cell and organismal biology is not fully appreciated. This has been the case due to the lack of the appropriate techniques and approaches. Heterogeneity can arise from alternative
[...] Read more.
Although reported in the literature, ribosome heterogeneity is a phenomenon whose extent and implications in cell and organismal biology is not fully appreciated. This has been the case due to the lack of the appropriate techniques and approaches. Heterogeneity can arise from alternative use and differential content of protein and RNA constituents, as well as from post-transcriptional and post-translational modifications. In the few examples we have, it is apparent that ribosomal heterogeneity offers an additional level and potential for gene expression regulation and might be a way towards tuning metabolism, stress, and growth programs to external and internal stimuli and needs. Here, we introduce ribosome biogenesis and discuss ribosomal heterogeneity in various reported occasions. We conclude that a systematic approach in multiple organisms will be needed to delineate this biological phenomenon and its contributions to growth, aging, and disease. Finally, we discuss ribosome mutations and their roles in disease.
Full article
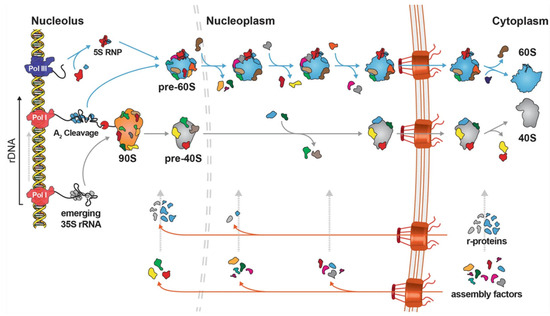
Figure 1
Open AccessReview
Evaluation of Exosomal miRNA as Potential Biomarkers in Cervical Cancer
by
, , , , , , and
Epigenomes 2023, 7(3), 16; https://doi.org/10.3390/epigenomes7030016 - 01 Aug 2023
Abstract
Different studies show that small non-coding RNAs, such as microRNAs (miRNAs) obtained from exosomes, are considered potential biomarkers in several types of cancer, including cervical cancer (CC). Therefore, the present study seeks to present an overview of the role of circulating exosomal miRNAs
[...] Read more.
Different studies show that small non-coding RNAs, such as microRNAs (miRNAs) obtained from exosomes, are considered potential biomarkers in several types of cancer, including cervical cancer (CC). Therefore, the present study seeks to present an overview of the role of circulating exosomal miRNAs with the potential to act as biomarkers for the diagnosis and prognosis of CC and to analyze the presence of these miRNAs according to the stage of CC. For this purpose, a review was developed, with articles consulted from the electronic databases MEDLINE/PubMed, Scopus, and Web of Science published between 2015 and 2021. Seven articles were included after a selection of studies according to the eligibility criteria. In addition to the methods used for sample analysis, detection, and isolation of miRNAs in each article, clinical data were also extracted from the patients studied, such as the stage of cancer. After analyzing the network of the seven miRNAs, they were associated with the immune system, CC progression and staging, and cisplatin resistance. With the belief that studies on miRNAs in cervical cancer would have major clinical implications, in this review, we have attempted to summarize the current situation and potential development prospects.
Full article
(This article belongs to the Special Issue Epigenetic Regulation in Cancer)
►▼
Show Figures

Figure 1
Open AccessArticle
Association of Toll-Like Receptor Gene Polymorphisms with Tuberculosis in HIV-Positive Participants
by
, , , , , and
Epigenomes 2023, 7(3), 15; https://doi.org/10.3390/epigenomes7030015 - 25 Jul 2023
Abstract
Genetic factors in the HIV-background may play a significant role in the susceptibility to secondary diseases, like tuberculosis, which is the leading cause in mortality of HIV-positive people. Toll-like receptors (TLRs) are considered to be receptors for adaptive immunity, and polymorphisms in TLR
[...] Read more.
Genetic factors in the HIV-background may play a significant role in the susceptibility to secondary diseases, like tuberculosis, which is the leading cause in mortality of HIV-positive people. Toll-like receptors (TLRs) are considered to be receptors for adaptive immunity, and polymorphisms in TLR genes can influence the activity of the immune response to infection. We conducted a case–control study of the association of TLR gene polymorphisms with the risk of tuberculosis coinfection in a multi-country sample of HIV-positive participants. Our study revealed certain associations between TLR4 and TLR6 polymorphisms and HIV–tuberculosis coinfection. We also found that the analyzed TLR1 and TLR4 polymorphisms were linked with the decline in CD4+ cell count, which is a predictor of disease progression in HIV-infected individuals. Our findings confirm that TLR gene polymorphisms are factors that may contribute to development of HIV–tuberculosis coinfection. However, the essence of the observed associations remains unclear, since it can also include both environmental factors and epigenetic mechanisms of gene expression regulation.
Full article
(This article belongs to the Topic Biosocial Studies)
►▼
Show Figures
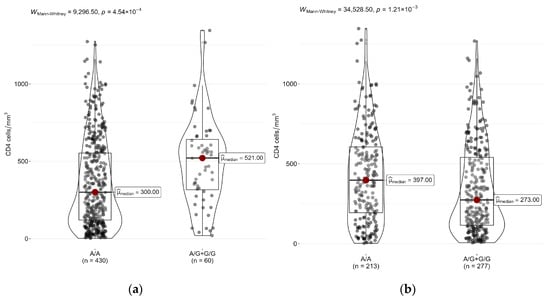
Figure 1

Journal Menu
► ▼ Journal Menu-
- Epigenomes Home
- Aims & Scope
- Editorial Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Topical Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics

Conferences
Special Issues
Special Issue in
Epigenomes
Epigenetic and Epitranscriptomic Determinants of Host-Microbe Interactions
Guest Editors: Mudasir Rashid, Peter C. Dedon, Manjula DasDeadline: 31 January 2024
Special Issue in
Epigenomes
RNA Modification in Inflammation and Metabolism
Guest Editors: Kezhong Zhang, Juncheng Wei, Cristina Espinosa-DiezDeadline: 30 April 2024
Special Issue in
Epigenomes
A Commemorative Issue in Honor of the 30th Anniversary of the Epigenetics Society
Guest Editors: Humaira Gowher, Assam El-Osta, Andrea FusoDeadline: 31 December 2024
Topical Collections
Topical Collection in
Epigenomes
Epigenetic Mechanisms in Diabetes Research
Collection Editor: Assam El-Osta
Topical Collection in
Epigenomes
Feature Papers in Epigenomes
Collection Editors: Ivana De la Serna, Che-Kun James Shen