Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
A Novel Bifidobacterium longum ssp. longum Strain with Pleiotropic Effects
Microorganisms 2024, 12(1), 174; https://doi.org/10.3390/microorganisms12010174 - 15 Jan 2024
Abstract
Postbiotics are gaining increasing interest among the scientific community as well as at the level of food processing enterprises. The aim of this preliminary study was to characterise the metabolic diversity of a novel Bifidobacterium longum strain, BIOCC 1719, of human origin. The
[...] Read more.
Postbiotics are gaining increasing interest among the scientific community as well as at the level of food processing enterprises. The aim of this preliminary study was to characterise the metabolic diversity of a novel Bifidobacterium longum strain, BIOCC 1719, of human origin. The change after 24 h cultivation in three media was assessed using a metabolomic approach. Milk-based substrates favoured the activity of the strain, promoting the production of B vitamins, essential amino acids, bile acids, and fatty acids. Vitamins B1, B2, B6, B7, and B12 (with an average increase of 20–30%) were produced in both whole milk and whey; the increased production in the latter was as high as 100% for B7 and 744% for B12. The essential amino acids methionine and threonine were produced (>38%) in both milk and whey, and there was an increased production of leucine (>50%) in milk and lysine (126%) in whey. Increases in the content of docosahexaenoic acid (DHA) by 20%, deoxycholic acid in milk and whey (141% and 122%, respectively), and cholic acid (52%) in milk were recorded. During the preliminary characterisation of the metabolic diversity of the novel B. longum strain, BIOCC 1719, we identified the bioactive compounds produced by the strain during fermentation. This suggests its potential use as a postbiotic ingredient to enrich the human diet.
Full article
(This article belongs to the Special Issue The Metabolism of Lactobacilli: Molecular Mechanisms and Applications)
Open AccessArticle
Monitoring of Animal Feed Contamination by Mycotoxins: Results of Five Years of Official Control by an Accredited Italian Laboratory
Microorganisms 2024, 12(1), 173; https://doi.org/10.3390/microorganisms12010173 - 15 Jan 2024
Abstract
Mycotoxin contamination of animal feed is a complex issue in both animal wellness and food safety. The most diffused mycotoxins subject to the official control of animal feed are Aflatoxin B1 (AF), Zearalenone (ZEA), Deoxynivalenol (DON), Ochratoxin A (OCRA), Fumonisins (FUMO), and T-2/HT-2
[...] Read more.
Mycotoxin contamination of animal feed is a complex issue in both animal wellness and food safety. The most diffused mycotoxins subject to the official control of animal feed are Aflatoxin B1 (AF), Zearalenone (ZEA), Deoxynivalenol (DON), Ochratoxin A (OCRA), Fumonisins (FUMO), and T-2/HT-2 toxins. This work describes the results of five years of monitoring focused on the evaluation of mycotoxin contamination of animal feed. Analytical determinations were carried out by means of accredited ELISA. The obtained results showed a non-alarming scenario, with several samples resulting as “non-compliant” according to the Maximum Residue Limits (MRLs) set in European Regulation No. 574/2011. Out of 722 analyzed samples coming from 2 Italian regions, Apulia and Basilicata, 14 samples were characterized by mycotoxin concentrations higher than related MRL; in particular, 5, 4, and 5 non-compliant samples for DON, AF, and ZEA, respectively. This study also evaluated the possible correlations between mycotoxin type and feed use with a special focus on animal sensitivity to mycotoxins.
Full article
(This article belongs to the Special Issue Mycotoxins in Food Safety, Food Security and Sustainability)
Open AccessArticle
Prevalence of Toxoplasma gondii IgG Antibodies and Associated Risk Factors in Psychiatric Patients from Western Romania: A Cross-Sectional Study
by
, , , , and
Microorganisms 2024, 12(1), 172; https://doi.org/10.3390/microorganisms12010172 - 15 Jan 2024
Abstract
Infection with the coccidian parasite Toxoplasma gondii was associated with an increased risk of several mental disorders. We conducted a case–control study of 464 consecutive psychiatric patients and assessed the prevalence of IgG antibodies against T. gondii and the potential risk factors associated
[...] Read more.
Infection with the coccidian parasite Toxoplasma gondii was associated with an increased risk of several mental disorders. We conducted a case–control study of 464 consecutive psychiatric patients and assessed the prevalence of IgG antibodies against T. gondii and the potential risk factors associated with infection. T. gondii-specific antibodies were determined using a chemiluminescence assay. A questionnaire was utilized to assess the potential correlation between risk factors and Toxoplasma gondii seropositivity. IgG antibodies were found in 325 (70.04%) of the patients. We observed a higher likelihood of positive IgG antibodies against Toxoplasma gondii in older individuals, patients residing in rural areas, and females. We also noted associations between Toxoplasma gondii infection and certain risk factors, like activities that involve contact with soil, low-income levels, and limited educational attainment. Our findings indicate a high prevalence of T. gondii infection among psychiatric patients from Western Romania and provide new information regarding the potential risk factors associated with T. gondii in this population group. This study may serve as a foundation for future research and the development of preventive strategies.
Full article
(This article belongs to the Special Issue State-of-the-Art Parasitic and Bacterial Infections in Romania)
Open AccessReview
Epidemiology of Cefotaxime-Hydrolysing β-Lactamase-Producing Escherichia coli in Children with Diarrhoea Reported Globally between 2012 and 2022
Microorganisms 2024, 12(1), 171; https://doi.org/10.3390/microorganisms12010171 - 15 Jan 2024
Abstract
The global spread of cefotaxime-hydrolysing β-lactamase (CTX-M)-producing Escherichia coli (E. coli) and its associated impact on paediatric diarrhoeal treatment and management has become a public health concern. This review assessed surveillance studies on CTX-M-producing E. coli associated with diarrhoea in children
[...] Read more.
The global spread of cefotaxime-hydrolysing β-lactamase (CTX-M)-producing Escherichia coli (E. coli) and its associated impact on paediatric diarrhoeal treatment and management has become a public health concern. This review assessed surveillance studies on CTX-M-producing E. coli associated with diarrhoea in children published between 2012 and 2022 globally. A total of thirty-eight studies were included for data analysis, categorised into continental regions, and tabulated. The majority (68%) of studies were conducted in Asian countries while few studies were conducted in Europe (11%) and Africa (18%), respectively. On the African continent, the majority (11%) of studies were conducted in Northern Africa while no studies were reported in East Africa. On the American continent, 3% of the studies were reported from South America. The studies included were classified into diarrheagenic E. coli (74%; 28/38) and faecal carriage (26%; 10/38). Of all the E. coli pathotypes associated with CTX-M production, EPEC was frequently reported. The prevalence of CTX-M-producing E. coli including the CTX-M-15-producing variants ranged between 1% and 94%. About 37% of the studies generalised the report as blaCTX-M-positive E. coli. The use of sequencing in characterising the CTX-M-producing E. coli was reported in only 32% of all the studies. This review provides information on the epidemiology of CTX-M-15-producing E. coli in paediatric diarrhoea and the extent to which surveillance is being performed. This is relevant in informing clinical practice for the management of diarrhoea as well as the design of future surveillance studies.
Full article
(This article belongs to the Special Issue Virulence Factors and Antibiotic Resistance of Enterobacterales 2.0)
►▼
Show Figures
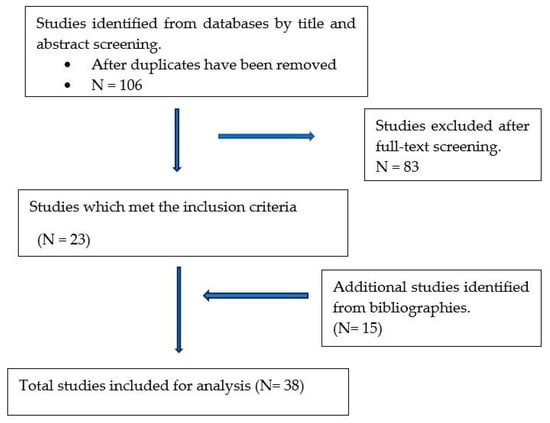
Figure 1
Open AccessArticle
An Analysis of the Gut Microbiota and Related Metabolites following PCSK9 Inhibition in Statin-Treated Patients with Elevated Levels of Lipoprotein(a)
by
, , , , , , , and
Microorganisms 2024, 12(1), 170; https://doi.org/10.3390/microorganisms12010170 - 15 Jan 2024
Abstract
Background. Atherosclerotic cardiovascular disease (ASCVD) is a leading cause of global mortality, often associated with high blood levels of LDL cholesterol (LDL-c). Medications like statins and PCSK9 inhibitors, are used to manage LDL-c levels and reduce ASCVD risk. Recent findings connect the gut
[...] Read more.
Background. Atherosclerotic cardiovascular disease (ASCVD) is a leading cause of global mortality, often associated with high blood levels of LDL cholesterol (LDL-c). Medications like statins and PCSK9 inhibitors, are used to manage LDL-c levels and reduce ASCVD risk. Recent findings connect the gut microbiota and its metabolites to ASCVD development. We showed that statins modulate the gut microbiota including the production of microbial metabolites involved in the regulation of cholesterol metabolism such as short chain fatty acids (SCFAs) and bile acids (BAs). Whether this pleiotropic effect of statins is associated with their antimicrobial properties or it is secondary to the modulation of cholesterol metabolism in the host is unknown. In this observational study, we evaluated whether alirocumab, a PCSK9 inhibitor administered subcutaneously, alters the stool-associated microbiota and the profiles of SCFAs and BAs. Methods. We used stool and plasma collected from patients enrolled in a single-sequence study using alirocumab. Microbial DNA was extracted from stool, and the bacterial component of the gut microbiota profiled following an amplicon sequencing strategy targeting the V3-V4 region of the 16S rRNA gene. Bile acids and SCFAs were profiled and quantified in stool and plasma using mass spectrometry. Results. Treatment with alirocumab did not alter bacterial alpha (Shannon index, p = 0.74) or beta diversity (PERMANOVA, p = 0.89) in feces. Similarly, circulating levels of SCFAs (mean difference (95% confidence interval (CI)), 8.12 [−7.15–23.36] µM, p = 0.25) and BAs (mean difference (95% CI), 0.04 [−0.11–0.19] log10(nmol mg−1 feces), p = 0.56) were equivalent regardless of PCSK9 inhibition. Alirocumab therapy was associated with increased concentration of BAs in feces (mean difference (95% CI), 0.20 [0.05–0.34] log10(nmol mg−1 feces), p = 0.01). Conclusion. In statin-treated patients, the use of alirocumab to inhibit PCSK9 leads to elevated levels of fecal BAs without altering the bacterial population of the gut microbiota. The association of alirocumab with increased fecal BA concentration suggests an additional mechanism for the cholesterol-lowering effect of PCSK9 inhibition.
Full article
(This article belongs to the Special Issue Gut Microbiome in Homeostasis and Disease 2.0)
►▼
Show Figures
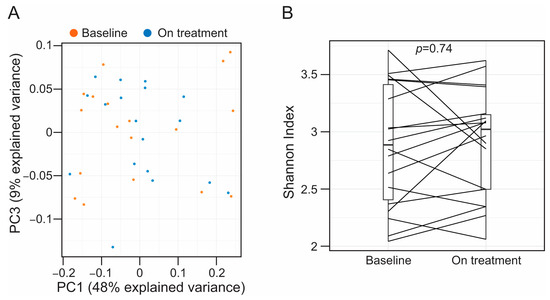
Figure 1
Open AccessArticle
Parenteral Vaccination with a Live Brucella melitensis Mutant Protects against Wild-Type B. melitensis 16M Challenge
Microorganisms 2024, 12(1), 169; https://doi.org/10.3390/microorganisms12010169 - 15 Jan 2024
Abstract
Susceptibility to brucellosis remains prevalent, even in herds vaccinated with conventional vaccines. Efforts are underway to develop an improved brucellosis vaccine, and possibly a universal vaccine, given that Brucella species are highly homologous. To this end, two B. melitensis mutants were developed, znBM-
[...] Read more.
Susceptibility to brucellosis remains prevalent, even in herds vaccinated with conventional vaccines. Efforts are underway to develop an improved brucellosis vaccine, and possibly a universal vaccine, given that Brucella species are highly homologous. To this end, two B. melitensis mutants were developed, znBM-lacZ (znBMZ) and znBM-mCherry (znBM-mC), and were tested for their ability to confer systemic immunity against virulent B. melitensis challenge. To assess the extent of their attenuation, bone-marrow-derived macrophages and human TF-1 myeloid cells were infected with both mutants, and the inability to replicate within these cells was noted. Mice infected with varying doses of znBM-mC cleared the brucellae within 6–10 weeks. To test for efficacy against systemic disease, groups of mice were vaccinated once by the intraperitoneal route with either znBMZ or B. abortus S19 vaccine. Relative to the PBS-dosed mice, znBMZ vaccination greatly reduced splenic brucellae colonization by ~25,000-fold compared to 700-fold for S19-vaccinated mice. Not surprisingly, both znBMZ and S19 strains induced IFN-γ+ CD4+ T cells, yet only znBMZ induced IFN-γ+ CD8+ T cells. While both strains induced CD4+ effector memory T cells (Tems), only znBMZ induced CD8+ Tems. Thus, these results show that the described znBM mutants are safe, able to elicit CD4+ and CD8+ T cell immunity without a boost, and highly effective, rendering them promising vaccine candidates for livestock.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►▼
Show Figures
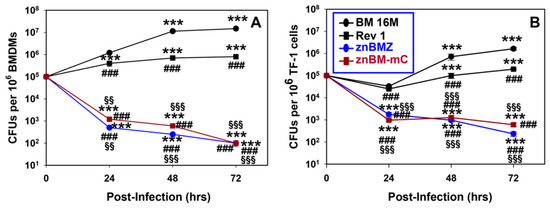
Figure 1
Open AccessReview
Potential for Use of Species in the Subfamily Erynioideae for Biological Control and Biotechnology
by
, , , , , , , , , and
Microorganisms 2024, 12(1), 168; https://doi.org/10.3390/microorganisms12010168 - 14 Jan 2024
Abstract
The fungal order Entomophthorales in the Zoopagomycota includes many fungal pathogens of arthropods. This review explores six genera in the subfamily Erynioideae within the family Entomophthoraceae, namely, Erynia, Furia, Orthomyces, Pandora, Strongwellsea, and Zoophthora. This is
[...] Read more.
The fungal order Entomophthorales in the Zoopagomycota includes many fungal pathogens of arthropods. This review explores six genera in the subfamily Erynioideae within the family Entomophthoraceae, namely, Erynia, Furia, Orthomyces, Pandora, Strongwellsea, and Zoophthora. This is the largest subfamily in the Entomophthorales, including 126 described species. The species diversity, global distribution, and host range of this subfamily are summarized. Relatively few taxa are geographically widespread, and few have broad host ranges, which contrasts with many species with single reports from one location and one host species. The insect orders infected by the greatest numbers of species are the Diptera and Hemiptera. Across the subfamily, relatively few species have been cultivated in vitro, and those that have require more specialized media than many other fungi. Given their potential to attack arthropods and their position in the fungal evolutionary tree, we discuss which species might be adopted for biological control purposes or biotechnological innovations. Current challenges in the implementation of these species in biotechnology include the limited ability or difficulty in culturing many in vitro, a correlated paucity of genomic resources, and considerations regarding the host ranges of different species.
Full article
(This article belongs to the Special Issue Advances in Research on Ancient Terrestrial Fungi)
►▼
Show Figures
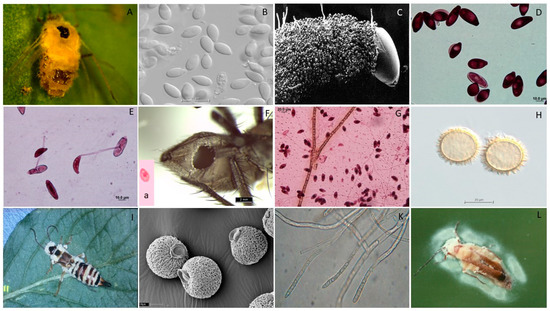
Figure 1
Open AccessArticle
Human Exposure to Naturally Occurring Bacillus anthracis in the Kars Region of Eastern Türkiye
by
, , , , , and
Microorganisms 2024, 12(1), 167; https://doi.org/10.3390/microorganisms12010167 - 14 Jan 2024
Abstract
Environmental contamination with Bacillus anthracis spores poses uncertain threats to human health. We undertook a study to determine whether inhabitants of the anthrax-endemic region of Kars in eastern Türkiye could develop immune responses to anthrax toxins without recognised clinical infection. We measured anti-PA
[...] Read more.
Environmental contamination with Bacillus anthracis spores poses uncertain threats to human health. We undertook a study to determine whether inhabitants of the anthrax-endemic region of Kars in eastern Türkiye could develop immune responses to anthrax toxins without recognised clinical infection. We measured anti-PA and anti-LF IgG antibody concentrations by ELISA in serum from 279 volunteers, 105 of whom had previously diagnosed anthrax infection (100 cutaneous, 5 gastrointestinal). Of the 174 without history of infection, 72 had prior contact with anthrax-contaminated material. Individuals were classified according to demographic parameters, daily working environment, and residence type. All villages in this study had recorded previous animal or human anthrax cases. Stepwise regression analyses showed that prior clinical infection correlated strongly with concentrations at the upper end of the ranges observed for both antibodies. For anti-PA, being a butcher and duration of continuous exposure risk correlated with high concentrations, while being a veterinarian or shepherd, time since infection, and town residence correlated with low concentrations. For anti-LF, village residence correlated with high concentrations, while infection limited to fingers or thumbs correlated with low concentrations. Linear discriminant analysis identified antibody concentration profiles associated with known prior infection. Profiles least typical of prior infection were observed in urban dwellers with known previous infection and in veterinarians without history of infection. Four individuals without history of infection (two butchers, two rural dwellers) had profiles suggesting unrecognised prior infection. Healthy humans therefore appear able to tolerate low-level exposure to environmental B. anthracis spores without ill effect, but it remains to be determined whether this exposure is protective. These findings have implications for authorities tasked with reducing the risk posed to human health by spore-contaminated materials and environments.
Full article
(This article belongs to the Special Issue Bacillus anthracis: Infection Mechanisms, Vaccination and Immune Response)
►▼
Show Figures
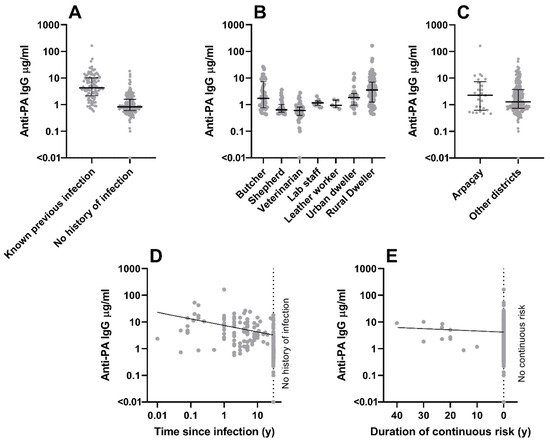
Figure 1
Open AccessReview
Valorization of Food Waste into Single-Cell Protein: An Innovative Technological Strategy for Sustainable Protein Production
Microorganisms 2024, 12(1), 166; https://doi.org/10.3390/microorganisms12010166 - 13 Jan 2024
Abstract
The rapidly increasing population and climate change pose a great threat to our current food systems. Moreover, the high usage of animal-based and plant-based protein has its drawbacks, as these nutritional sources require many hectares of land and water, are affected by seasonal
[...] Read more.
The rapidly increasing population and climate change pose a great threat to our current food systems. Moreover, the high usage of animal-based and plant-based protein has its drawbacks, as these nutritional sources require many hectares of land and water, are affected by seasonal variations, are costly, and contribute to environmental pollution. Single-cell proteins (SCPs) are gaining a lot of research interest due to their remarkable properties, such as their high protein content that is comparable with other protein sources; low requirements for land and water; low carbon footprint; and short production period. This review explores the use of food waste as a sustainable feedstock for the advancement of SCP processes. It discusses SCP studies that exploit food waste as a substrate, alongside the biocatalysts (bacteria, fungi, yeast, and microalgae) that are used. The operational setpoint conditions governing SCP yields and SCP fermentation routes are elucidated as well. This review also demonstrates how the biorefinery concept is implemented in the literature to improve the economic potential of “waste-to-protein” innovations, as this leads to the establishment of multiproduct value chains. A short section that discusses the South African SCP scenario is also included. The technical and economic hurdles facing second-generation SCP processes are also discussed, together with future perspectives. Therefore, SCP technologies could play a crucial role in the acceleration of a “sustainable protein market”, and in tackling the global hunger crisis.
Full article
(This article belongs to the Special Issue The Role of Microbes in Biorefinery Products and Biofuels)
►▼
Show Figures
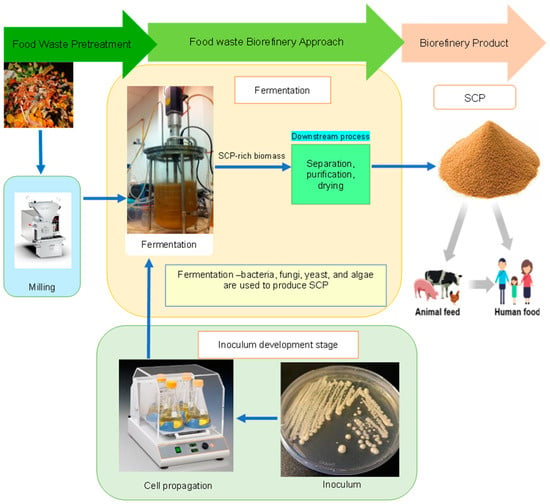
Figure 1
Open AccessCase Report
Fowl Typhoid Outbreak on a Commercial Turkey Farm in Croatia
Microorganisms 2024, 12(1), 165; https://doi.org/10.3390/microorganisms12010165 - 13 Jan 2024
Abstract
Fowl typhoid is a septicemic disease caused by Salmonella enterica subsp. enterica serovar Gallinarum biovar Gallinarum. It is a host-specific disease primarily affecting chickens and turkeys, although it has been reported in various animal species and sporadically in humans. Here, we present a
[...] Read more.
Fowl typhoid is a septicemic disease caused by Salmonella enterica subsp. enterica serovar Gallinarum biovar Gallinarum. It is a host-specific disease primarily affecting chickens and turkeys, although it has been reported in various animal species and sporadically in humans. Here, we present a case of a fowl typhoid outbreak on a turkey poult farm where the source of infection was the hatchery. The birds started showing symptoms of growth retardation at 21 days of age, after which the mortality rates gradually started to increase. Post mortem examination revealed that the main lesions were granulomatous proliferations in the small intestines. The results of the histopathological examination indicate that the severity of the infection was alleviated by the application of phytogenic mixtures and probiotics as a supportive treatment, even though the affected flock was eventually culled at 60 days of age. The farmer was advised to apply more strict biosecurity measures to prevent the spread of the disease on the farm and try to eradicate the pathogen from the barn. Since the outbreak, there have been no recurrent infections.
Full article
(This article belongs to the Special Issue Avian Pathogens 2.0)
►▼
Show Figures
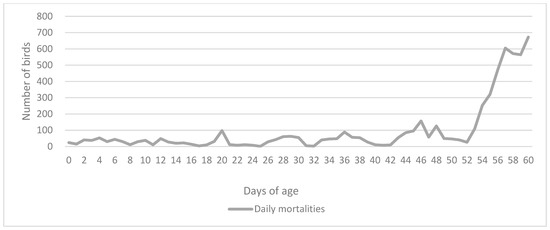
Figure 1
Open AccessReview
Infective Endocarditis Due to Kingella kingae
by
, , , , , , , , and
Microorganisms 2024, 12(1), 164; https://doi.org/10.3390/microorganisms12010164 - 13 Jan 2024
Abstract
Infective endocarditis due to Kingella kingae is a rare but serious invasive infection that occurs mostly in children. Recent advances in nucleic acid amplification testing as well as in cardiac imaging have enabled more accurate diagnosis. A good understanding of the epidemiology and
[...] Read more.
Infective endocarditis due to Kingella kingae is a rare but serious invasive infection that occurs mostly in children. Recent advances in nucleic acid amplification testing as well as in cardiac imaging have enabled more accurate diagnosis. A good understanding of the epidemiology and virulence factors remains crucial to guide the therapeutic approach. Here, we synthesize the current state of knowledge on epidemiological features, pathophysiological insights, complications, and therapy regarding Kingella kingae endocarditis in children and adults. Finally, throughout this comprehensive review, knowledge gaps and areas for future research are also identified.
Full article
(This article belongs to the Special Issue Kingella kingae: Virulence Factors, Clinical Disease, and Diagnostics)
►▼
Show Figures
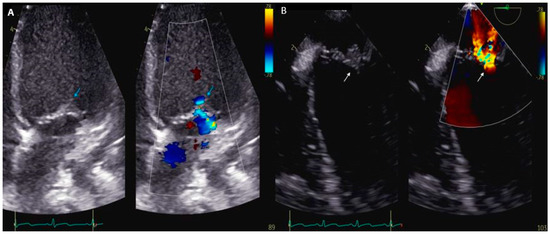
Figure 1
Open AccessEditorial
Soil Fungi in Sustainable Agriculture
Microorganisms 2024, 12(1), 163; https://doi.org/10.3390/microorganisms12010163 - 13 Jan 2024
Abstract
It is widely accepted that the continuously growing human population needs rapid solutions to respond to the increased global demand for high agricultural productivity [...]
Full article
(This article belongs to the Special Issue Soil Fungi in Sustainable Agriculture)
Open AccessArticle
Comparison of Experimental Methodologies Based on Bulk-Metagenome and Virus-like Particle Enrichment: Pros and Cons for Representativeness and Reproducibility in the Study of the Fecal Human Virome
by
, , , , and
Microorganisms 2024, 12(1), 162; https://doi.org/10.3390/microorganisms12010162 - 13 Jan 2024
Abstract
Studies on the human virome based on the application of metagenomic approaches involve overcoming a series of challenges and limitations inherent not only to the biological features of viruses, but also to methodological pitfalls which different approaches have tried to minimize. These approaches
[...] Read more.
Studies on the human virome based on the application of metagenomic approaches involve overcoming a series of challenges and limitations inherent not only to the biological features of viruses, but also to methodological pitfalls which different approaches have tried to minimize. These approaches fall into two main categories: bulk-metagenomes and virus-like particle (VLP) enrichment. In order to address issues associated with commonly used experimental procedures to assess the degree of reliability, representativeness, and reproducibility, we designed a comparative analysis applied to three experimental protocols, one based on bulk-metagenomes and two based on VLP enrichment. These protocols were applied to stool samples from 10 adult participants, including two replicas per protocol and subject. We evaluated the performances of the three methods, not only through the analysis of the resulting composition, abundance, and diversity of the virome via taxonomical classification and type of molecule (DNA versus RNA, single stranded vs. double stranded), but also according to how the a priori identical replicas differed from each other according to the extraction methods used. Our results highlight the strengths and weaknesses of each approach, offering valuable insights and tailored recommendations for drawing reliable conclusions based on specific research goals.
Full article
(This article belongs to the Special Issue Advances in Viral Metagenomics)
►▼
Show Figures
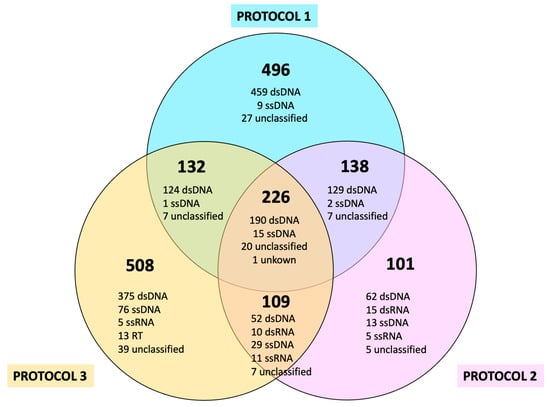
Figure 1
Open AccessReview
Recent Advances in Diagnosis and Treatment Approaches in Fungal Keratitis: A Narrative Review
by
, , , and
Microorganisms 2024, 12(1), 161; https://doi.org/10.3390/microorganisms12010161 - 13 Jan 2024
Abstract
Fungal keratitis represents a potentially sight-threatening infection associated with poor prognosis, as well as financial burden. Novel diagnostic methods include polymerase-chain-reaction (PCR)-based approaches, metagenomic deep sequences, in vivo confocal microscopy, and antifungal susceptibility testing. The ideal therapeutic approaches and outcomes have been widely
[...] Read more.
Fungal keratitis represents a potentially sight-threatening infection associated with poor prognosis, as well as financial burden. Novel diagnostic methods include polymerase-chain-reaction (PCR)-based approaches, metagenomic deep sequences, in vivo confocal microscopy, and antifungal susceptibility testing. The ideal therapeutic approaches and outcomes have been widely discussed in recent times, with early therapy being of the utmost importance for the preservation of visual acuity, minimizing corneal damage and reducing the scar size. However, combination therapy can be more efficacious compared to monotherapy. Understanding the pathogenesis, early diagnosis, and prevention strategies can be of great importance. In this narrative, we discuss the recent progress that may aid our understanding of the diagnosis, treatment, and prevention of mycotic keratitis.
Full article
(This article belongs to the Special Issue Infectious Eye Diseases and Prevention Control)
Open AccessArticle
Metabarcoding Analysis of Microorganisms Inside Household Washing Machines in Shanghai, China
by
, , , , , , , and
Microorganisms 2024, 12(1), 160; https://doi.org/10.3390/microorganisms12010160 - 13 Jan 2024
Abstract
Washing machines are one of the tools that bring great convenience to people’s daily lives. However, washing machines that have been used for a long time often develop issues such as odor and mold, which can pose health hazards to consumers. There exists
[...] Read more.
Washing machines are one of the tools that bring great convenience to people’s daily lives. However, washing machines that have been used for a long time often develop issues such as odor and mold, which can pose health hazards to consumers. There exists a conspicuous gap in our understanding of the microorganisms that inhabit the inner workings of washing machines. In this study, samples were collected from 22 washing machines in Shanghai, China, including both water eluted from different parts of washing machines and biofilms. Quantitative qualitative analysis was performed using fluorescence PCR quantification, and microbial communities were characterized by high-throughput sequencing (HTS). This showed that the microbial communities in all samples were predominantly composed of bacteria. HTS results showed that in the eluted water samples, the bacteria mainly included Pseudomonas, Enhydrobacter, Brevibacterium, and Acinetobacter. Conversely, in the biofilm samples, Enhydrobacter and Brevibacterium were the predominant bacterial microorganisms. Correlation analysis results revealed that microbial colonies in washing machines were significantly correlated with years of use and the type of detergent used to clean the washing machine. As numerous pathogenic microorganisms can be observed in the results, effective preventive measures and future research are essential to mitigate these health problems and ensure the continued safe use of these household appliances.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures
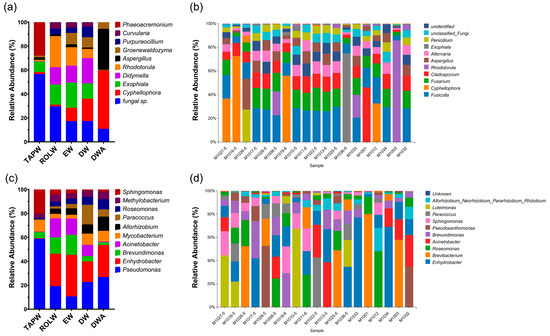
Figure 1
Open AccessCommunication
Whole Genome Sequences, De Novo Assembly, and Annotation of Antibiotic Resistant Campylobacter jejuni Strains S27, S33, and S36 Newly Isolated from Chicken Meat
Microorganisms 2024, 12(1), 159; https://doi.org/10.3390/microorganisms12010159 - 13 Jan 2024
Abstract
Campylobacter is a leading bacterial cause of gastrointestinal infections in humans and has imposed substantial medical and public health burdens worldwide. Among a total of 39 species in the Campylobacter genus, C. jejuni is the most important species responsible for approx. 90% of
[...] Read more.
Campylobacter is a leading bacterial cause of gastrointestinal infections in humans and has imposed substantial medical and public health burdens worldwide. Among a total of 39 species in the Campylobacter genus, C. jejuni is the most important species responsible for approx. 90% of human Campylobacter illness. Most cases of the infection were acquired by ingesting undercooked poultry meat due to the high prevalence of Campylobacter in the products. Here, we reported the dataset of raw sequences, de novo assembled and annotated genomes of C. jejuni strains S27, S33, and S36 recently isolated from retail chicken by using PacBio highly accurate long-read sequencing technology combined with bioinformatics tools. Our data revealed several virulence and antibiotic resistance genes in each of the chromosomes, a type IV secretion system in the plasmid (pCjS33) of C. jejuni S33, and a type IV secretion system and a phage in the plasmid (pCjS36) of C. jejuni S36. This study not only provides new sequence data but also extends the knowledge pertaining to the genomic and functional aspects of this important foodborne pathogen, including the genetic determinants of virulence and antibiotic resistance.
Full article
(This article belongs to the Special Issue Advances in Campylobacter: Molecular Epidemiology, Virulence Factors, Immune Response and Drug Resistance)
Open AccessCorrection
Correction: Mao et al. Gut Bacterial Community Determines the Therapeutic Effect of Ginsenoside on Canine Inflammatory Bowel Disease by Modulating the Colonic Mucosal Barrier. Microorganisms 2023, 11, 2616
Microorganisms 2024, 12(1), 158; https://doi.org/10.3390/microorganisms12010158 - 12 Jan 2024
Abstract
The authors wish to make the following corrections to this paper [...]
Full article
(This article belongs to the Special Issue The Role of Gut Microbiota in Inflammatory Bowel Disease: Mechanisms and Novel Therapeutic Approaches)
Open AccessArticle
Rapid Visual Detection of Methicillin-Resistant Staphylococcus aureus in Human Clinical Samples via Closed LAMP Assay Targeting mecA and spa Genes
by
, , , , , , and
Microorganisms 2024, 12(1), 157; https://doi.org/10.3390/microorganisms12010157 - 12 Jan 2024
Abstract
The emergence of antimicrobial resistance (AMR), particularly methicillin-resistant Staphylococcus aureus (MRSA), poses a significant global health threat as these bacteria increasingly become resistant to the most available therapeutic options. Thus, developing an efficient approach to rapidly screen MRSA directly from clinical specimens has
[...] Read more.
The emergence of antimicrobial resistance (AMR), particularly methicillin-resistant Staphylococcus aureus (MRSA), poses a significant global health threat as these bacteria increasingly become resistant to the most available therapeutic options. Thus, developing an efficient approach to rapidly screen MRSA directly from clinical specimens has become vital. In this study, we establish a closed-tube loop-mediated isothermal amplification (LAMP) method incorporating hydroxy-naphthol blue (HNB) colorimetric dye assay to directly detect MRSA from clinical samples based on the presence of mecA and spa genes. In total, 125 preidentified S. aureus isolates and 93 clinical samples containing S. aureus were sourced from the microbiology laboratory at Hamad General Hospital (HGH). The sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were computed based on conventional PCR. The assay demonstrated 100% specificity, 91.23% sensitivity, 0.90 Cohen Kappa (CK), 100% PPV, and 87.8% NPV for the clinical samples, while clinical isolates exhibited 100% specificity, 97% sensitivity, 0.926 CK, 100% PPV, and 88.89% NPV. Compared to cefoxitin disk diffusion, LAMP provided 100% specificity and sensitivity, 1.00 CK, and 100% for PPV and NPV. The study revealed that the closed-tube LAMP incorporating (HNB) dye is a rapid technique with a turnaround time of less than 1 h and high specificity and sensitivity.
Full article
(This article belongs to the Section Microbial Biotechnology)
►▼
Show Figures

Figure 1
Open AccessEditorial
An Open View on SARS-CoV-2 Infection
by
and
Microorganisms 2024, 12(1), 156; https://doi.org/10.3390/microorganisms12010156 - 12 Jan 2024
Abstract
The onset of the SARS-CoV-2 virus led to the appearance of a devastating pandemic, which once again demonstrated the practical importance of virology [...]
Full article
(This article belongs to the Special Issue SARS-CoV-2/COVID-19 Infection: Molecular and Clinical Aspects)
Open AccessReview
The Impact and Effects of Host Immunogenetics on Infectious Disease Studies Using Non-Human Primates in Biomedical Research
Microorganisms 2024, 12(1), 155; https://doi.org/10.3390/microorganisms12010155 - 12 Jan 2024
Abstract
Understanding infectious disease pathogenesis and evaluating novel candidate treatment interventions for human use frequently requires prior or parallel analysis in animal model systems. While rodent species are frequently applied in such studies, there are situations where non-human primate (NHP) species are advantageous or
[...] Read more.
Understanding infectious disease pathogenesis and evaluating novel candidate treatment interventions for human use frequently requires prior or parallel analysis in animal model systems. While rodent species are frequently applied in such studies, there are situations where non-human primate (NHP) species are advantageous or required. These include studies of animals that are anatomically more akin to humans, where there is a need to interrogate the complexity of more advanced biological systems or simply reflect susceptibility to a specific infectious agent. The contribution of different arms of the immune response may be addressed in a variety of NHP species or subspecies in specific physiological compartments. Such studies provide insights into immune repertoires not always possible from human studies. However, genetic variation in outbred NHP models may confound, or significantly impact the outcome of a particular study. Thus, host factors need to be considered when undertaking such studies. Considerable knowledge of the impact of host immunogenetics on infection dynamics was elucidated from HIV/SIV research. NHP models are now important for studies of emerging infections. They have contributed to delineating the pathogenesis of SARS-CoV-2/COVID-19, which identified differences in outcomes attributable to the selected NHP host. Moreover, their use was crucial in evaluating the immunogenicity and efficacy of vaccines against COVID-19 and establishing putative correlates of vaccine protection. More broadly, neglected or highly pathogenic emerging or re-emergent viruses may be studied in selected NHPs. These studies characterise protective immune responses following infection or the administration of candidate immunogens which may be central to the accelerated licensing of new vaccines. Here, we review selected aspects of host immunogenetics, specifically MHC background and TRIM5 polymorphism as exemplars of adaptive and innate immunity, in commonly used Old and New World host species. Understanding this variation within and between NHP species will ensure that this valuable laboratory source is used most effectively to combat established and emerging virus infections and improve human health worldwide.
Full article
(This article belongs to the Special Issue Emerging Infectious Diseases in Humans and Animals)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomass, Energies, Microorganisms, Processes, Water
Anaerobic Digestion Processes, 2nd Volume
Topic Editors: Yue Zhang, Davide DionisiDeadline: 31 January 2024
Topic in
Biology, Diversity, Microorganisms, Molecules, Life
Extreme Environments: Microbial and Biochemical Diversity
Topic Editors: Massimiliano Fenice, Susanna Gorrasi, Marcella PasqualettiDeadline: 31 March 2024
Topic in
Biology, Ecologies, Forests, Microorganisms, Plants
Litter Decompositions: From Individuals to Ecosystems
Topic Editors: Wen Zhou, Guihua LiuDeadline: 30 April 2024
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Emerging Research on Tick-Borne Pathogens and Diseases
Guest Editors: Leona Gilbert, John LambertDeadline: 20 January 2024
Special Issue in
Microorganisms
Assembly, Structure, and Germination of Bacterial Spores
Guest Editors: Graham Christie, Bing Hao, Peter SetlowDeadline: 30 January 2024
Special Issue in
Microorganisms
Vector-Borne Infections in Wildlife
Guest Editor: Marcos Rogério AndréDeadline: 15 February 2024
Special Issue in
Microorganisms
Microbiome in Infectious Diseases
Guest Editors: Elda Righi, Assunta SartorDeadline: 29 February 2024
Topical Collections
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu
Topical Collection in
Microorganisms
Microbial Life in Extreme Environments
Collection Editor: Ricardo Amils