Journal Description
Methods and Protocols
Methods and Protocols
is an international, peer-reviewed, open access journal aiming to establish and describe new experimental techniques in the fields of Life Sciences, Chemistry, and Biomedical Sciences, published bimonthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, CAPlus / SciFinder, and other databases.
- Journal Rank: CiteScore - Q2 (Biochemistry, Genetics and Molecular Biology (miscellaneous))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 27.9 days after submission; acceptance to publication is undertaken in 3.9 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
2.4 (2022);
5-Year Impact Factor:
2.4 (2022)
Latest Articles
Making the Most of Lateral Flow Immunochromatographic Tests: An Efficient Protocol to Recover DNA
Methods Protoc. 2024, 7(1), 8; https://doi.org/10.3390/mps7010008 - 15 Jan 2024
Abstract
Lateral flow immunochromatographic (LFI) tests are widely used in both biomedical and forensic sciences for different applications. In forensic sciences, their main use is to detect body fluids at crime scenes. However, there are situations in which the amount of potential biological evidence
[...] Read more.
Lateral flow immunochromatographic (LFI) tests are widely used in both biomedical and forensic sciences for different applications. In forensic sciences, their main use is to detect body fluids at crime scenes. However, there are situations in which the amount of potential biological evidence is so low that DNA extraction is favored with respect to the identification of body fluids. Here, an efficient and quick protocol is presented to integrate the detection of body fluids through LFI with DNA extraction from a sample swab and buffer, providing a complete characterization of the biological evidence. This protocol is a modification of a general DNA extraction silica-based kit, whose main application is for blood and tissues. Thus, it could be carried out in different settings (forensic labs, hospitals, other testing labs) without the necessity of buying a specific kit for swabs. The validation of this protocol is supported by the results presented here and previous publications from our group, obtaining DNA in good quantity and with good quality. This proves the potential application of the protocol in both forensic scenarios, to fully characterize biological evidence, and biomedical settings, to molecularly confirm the results of LFI tests.
Full article
(This article belongs to the Section Molecular and Cellular Biology)
►
Show Figures
Open AccessArticle
Mapping m6A Sites on HIV-1 RNA Using Oligonucleotide LC-MS/MS
by
, , , , , , , and
Ga-Eun Lee
Methods Protoc. 2024, 7(1), 7; https://doi.org/10.3390/mps7010007 - 10 Jan 2024
Abstract
The biological significance of chemical modifications to the ribonucleic acid (RNA) of human immunodeficiency virus type-1 (HIV-1) has been recognized. However, our understanding of the site-specific and context-dependent roles of these chemical modifications remains limited, primarily due to the absence of nucleotide-resolution mapping
[...] Read more.
The biological significance of chemical modifications to the ribonucleic acid (RNA) of human immunodeficiency virus type-1 (HIV-1) has been recognized. However, our understanding of the site-specific and context-dependent roles of these chemical modifications remains limited, primarily due to the absence of nucleotide-resolution mapping of modification sites. In this study, we present a method for achieving nucleotide-resolution mapping of chemical modification sites on HIV-1 RNA using liquid chromatography and tandem mass spectrometry (LC–MS/MS). LC–MS/MS, a powerful tool capable of directly analyzing native RNAs, has proven effective for mapping RNA modifications in small RNA molecules, including ribosomal RNA and transfer RNA. However, longer RNAs have posed challenges, such as the 9 Kb HIV-1 virion RNA, due to the complexity of and ambiguity in mass differences among RNase T1-cleaved RNA fragments in LC-MS/MS data. Here, we introduce a new target RNA enrichment method to isolate small local RNA fragments of HIV-1 RNA that potentially harbor site-specific N6-methyladenosine (m6A) modifications. In our initial trial, we used target-specific DNA probes only and encountered insufficient RNA fragmentation due to inefficient S1 digestion near the target site. Recognizing that inefficient S1 digestion by HIV-1 RNA is likely due to the formation of secondary structures in proximity to the target site, we designed multiple DNA probes annealing to various sites of HIV-1 RNA to better control the structures of RNA substrates for S1 digestion. The use of these non-target DNA probes significantly improved the isolation of more homogeneous target RNA fragments of approximately 50 bases in length. Oligonucleotide LC-MS/MS analysis of these isolated target RNA fragments successfully separated and detected both m6A-methylated and non-methylated oligomers at the two m6A-predicted sites. The principle of this new target enrichment strategy holds promise and should be broadly applicable to the analysis of any lengthy RNA that was previously deemed infeasible for investigation using oligonucleotide LC-MS/MS.
Full article
(This article belongs to the Section Biochemical and Chemical Analysis & Synthesis)
►▼
Show Figures
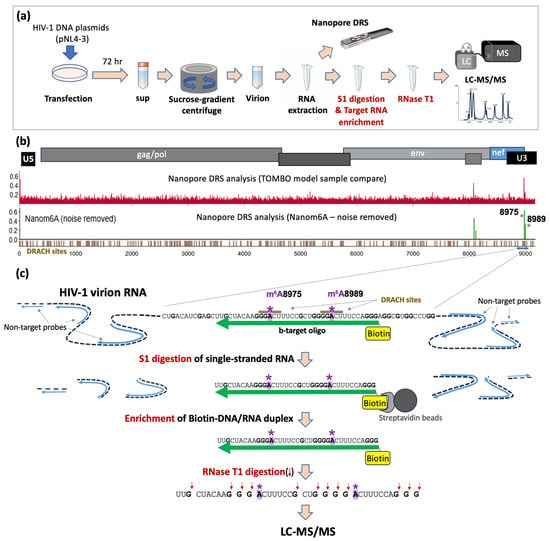
Figure 1
Open AccessArticle
Optimal Selection of Sampling Points within Sewer Networks for Wastewater-Based Epidemiology Applications
Methods Protoc. 2024, 7(1), 6; https://doi.org/10.3390/mps7010006 - 05 Jan 2024
Abstract
Wastewater-based epidemiology (WBE) has great potential to monitor community public health, especially during pandemics. However, it faces substantial hurdles in pathogen surveillance through WBE, encompassing data representativeness, spatiotemporal variability, population estimates, pathogen decay, and environmental factors. This paper aims to enhance the reliability
[...] Read more.
Wastewater-based epidemiology (WBE) has great potential to monitor community public health, especially during pandemics. However, it faces substantial hurdles in pathogen surveillance through WBE, encompassing data representativeness, spatiotemporal variability, population estimates, pathogen decay, and environmental factors. This paper aims to enhance the reliability of WBE data, especially for early outbreak detection and improved sampling strategies within sewer networks. The tool implemented in this paper combines a monitoring model and an optimization model to facilitate the optimal selection of sampling points within sewer networks. The monitoring model utilizes parameters such as feces density and average water consumption to define the detectability of the virus that needs to be monitored. This allows for standardization and simplicity in the process of moving from the analysis of wastewater samples to the identification of infection in the source area. The entropy-based model can select optimal sampling points in a sewer network to obtain the most specific information at a minimum cost. The practicality of our tool is validated using data from Hildesheim, Germany, employing SARS-CoV-2 as a pilot pathogen. It is important to note that the tool’s versatility empowers its extension to monitor other pathogens in the future.
Full article
(This article belongs to the Section Public Health Research)
►▼
Show Figures
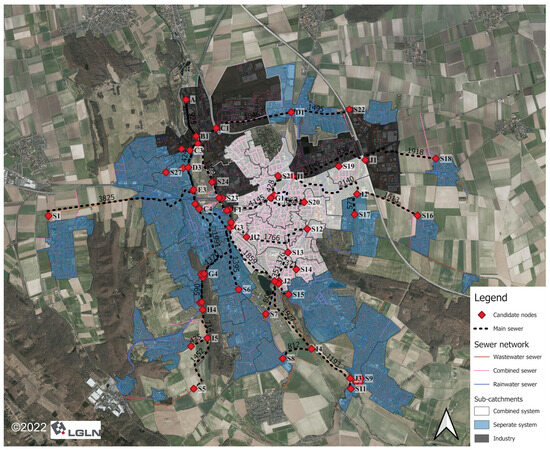
Figure 1
Open AccessStudy Protocol
Introducing Artificial Intelligence in Interpretation of Foetal Cardiotocography: Medical Dataset Curation and Preliminary Coding—An Interdisciplinary Project
by
, , , , , , , , and
Methods Protoc. 2024, 7(1), 5; https://doi.org/10.3390/mps7010005 - 04 Jan 2024
Abstract
Artificial intelligence (AI) is gaining increasing interest in the field of medicine because of its capacity to process big data and pattern recognition. Cardiotocography (CTG) is widely used for the assessment of foetal well-being and uterine contractions during pregnancy and labour. It is
[...] Read more.
Artificial intelligence (AI) is gaining increasing interest in the field of medicine because of its capacity to process big data and pattern recognition. Cardiotocography (CTG) is widely used for the assessment of foetal well-being and uterine contractions during pregnancy and labour. It is characterised by inter- and intraobserver variability in interpretation, which depends on the observers’ experience. Artificial intelligence (AI)-assisted interpretation could improve its quality and, thus, intrapartal care. Cardiotocography (CTG) raw signals from labouring women were extracted from the database at the University Hospital of Bern between 2006 and 2019. Later, they were matched with the corresponding foetal outcomes, namely arterial umbilical cord pH and 5-min APGAR score. Excluded were deliveries where data were incomplete, as well as multiple births. Clinical data were grouped regarding foetal pH and APGAR score at 5 min after delivery. Physiological foetal pH was defined as 7.15 and above, and a 5-min APGAR score was considered physiologic when reaching ≥7. With these groups, the algorithm was trained to predict foetal hypoxia. Raw data from 19,399 CTG recordings could be exported. This was accomplished by manually searching the patient’s identification numbers (PIDs) and extracting the corresponding raw data from each episode. For some patients, only one episode per pregnancy could be found, whereas for others, up to ten episodes were available. Initially, 3400 corresponding clinical outcomes were found for the 19,399 CTGs (17.52%). Due to the small size, this dataset was rejected, and a new search strategy was elaborated. After further matching and curation, 6141 (31.65%) paired data samples could be extracted (cardiotocography raw data and corresponding maternal and foetal outcomes). Of these, half will be used to train artificial intelligence (AI) algorithms, whereas the other half will be used for analysis of efficacy. Complete data could only be found for one-third of the available population. Yet, to our knowledge, this is the most exhaustive and second-largest cardiotocography database worldwide, which can be used for computer analysis and programming. A further enrichment of the database is planned.
Full article
(This article belongs to the Section Biomedical Sciences and Physiology)
►▼
Show Figures
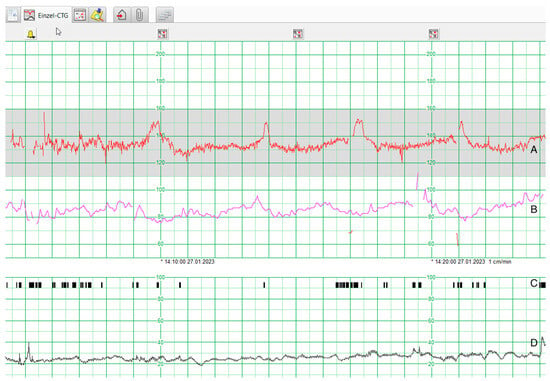
Figure 1
Open AccessStudy Protocol
A Scoping Review Protocol of Social Determinants of HIV/TB Coinfections in Sub-Saharan Africa
by
and
Methods Protoc. 2024, 7(1), 4; https://doi.org/10.3390/mps7010004 - 04 Jan 2024
Abstract
►▼
Show Figures
Introduction: Tuberculosis (TB) and Human Immunodeficiency Virus (HIV) remain major public health issues in sub-Saharan Africa. The co-occurrence of these diseases is a growing concern in the region, and social determinants, the circumstances under which people are born, live, work, and age, are
[...] Read more.
Introduction: Tuberculosis (TB) and Human Immunodeficiency Virus (HIV) remain major public health issues in sub-Saharan Africa. The co-occurrence of these diseases is a growing concern in the region, and social determinants, the circumstances under which people are born, live, work, and age, are known to influence the risk of disease transmission, diagnosis, treatment, and outcomes. Here, we present a protocol for the evidence synthesis on the social determinants of HIV/TB coinfections in sub-Saharan Africa. The high prevalence of Tuberculosis (TB) and Human Immunodeficiency Virus (HIV) in sub-Saharan Africa presents significant public health challenges. TB/HIV comorbidity is influenced by various social determinants, including social, economic, cultural, and environmental factors, impacting disease transmission risk, accurate diagnosis, and treatment outcomes. This study protocol aims to provide an evidence synthesis on the social determinants of HIV/TB coinfection in sub-Saharan Africa. Methods and analysis: The researchers will use the Arksey and O’Malley’s (2005) methodological framework to guide the scoping review. First, databases such as PubMed, MEDLINE, Web of Science, and PsychInfo will be searched. The researchers will then proceed in two steps. Before finalising the study selection, two independent reviewers will examine the article titles and abstracts for eligibility and inclusion. The researchers will then conduct a full-text screening of the articles based on the selected titles and abstracts. The authors’ tool will be used to extract data, ensuring that the articles are properly screened and that the risk of bias is minimized. The chosen studies will be examined using a standardized tool to examine all bibliographic data and study characteristics. Ethics and dissemination: The review will provide an overview of the social determinants influencing the prevalence and outcomes of TB/HIV comorbidity in the region, as well as identify any research gaps. Policymakers, researchers, and healthcare professionals will benefit from the findings in developing targeted interventions to address the social determinants of TB/HIV comorbidity in sub-Saharan Africa.
Full article
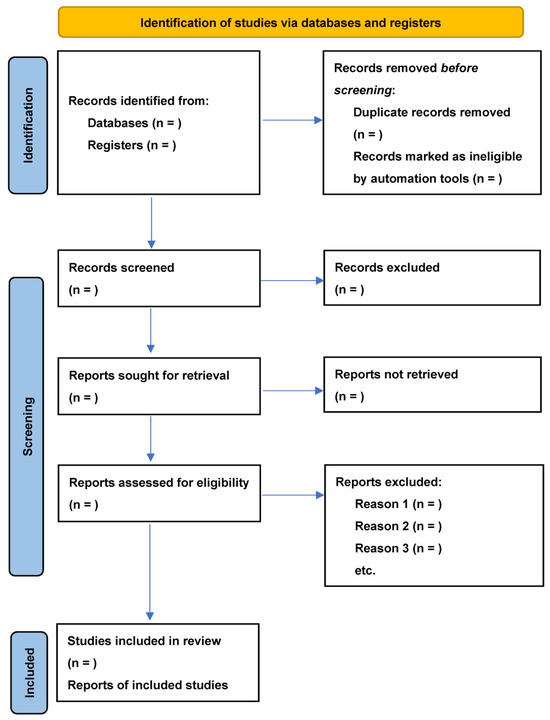
Figure 1
Open AccessProtocol
Generation of Viral Particles with Brain Cell-Specific Tropism by Pseudotyping HIV-1 with the Zika Virus E Protein
Methods Protoc. 2024, 7(1), 3; https://doi.org/10.3390/mps7010003 - 28 Dec 2023
Abstract
Flaviviruses are a family of RNA viruses that includes many known pathogens, such as Zika virus (ZIKV), West Nile virus (WNV), dengue virus (DENV), and yellow fever virus (YFV). A pseudotype is an artificial virus particle created in vitro by incorporating the flavivirus
[...] Read more.
Flaviviruses are a family of RNA viruses that includes many known pathogens, such as Zika virus (ZIKV), West Nile virus (WNV), dengue virus (DENV), and yellow fever virus (YFV). A pseudotype is an artificial virus particle created in vitro by incorporating the flavivirus envelope proteins into the structure of, for example, a retrovirus such as human immunodeficiency virus type-1 (HIV-1). They can be a useful tool in virology for understanding the biology of flaviviruses, evaluating immune responses, developing antiviral strategies but can also be used as vectors for gene transfer experiments. This protocol describes the generation of a ZIKV/HIV-1 pseudotype developed as a new tool for infecting cells derived from a highly malignant brain tumor: glioblastoma multiforme grade 4.
Full article
(This article belongs to the Section Molecular and Cellular Biology)
►▼
Show Figures
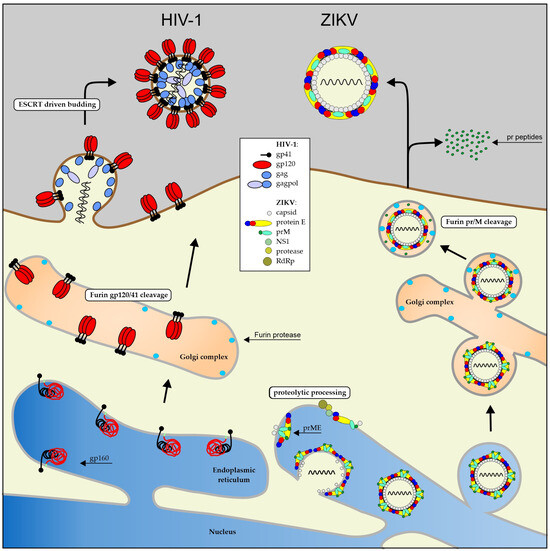
Figure 1
Open AccessArticle
Nesting Ecology of Lepidochelys olivacea in Lobito, Angola
by
, , , , and
Methods Protoc. 2024, 7(1), 2; https://doi.org/10.3390/mps7010002 - 27 Dec 2023
Abstract
►▼
Show Figures
The scarcity on the Atlantic coast of the African sea turtle population and its dynamics data is well known. This article discusses the nesting ecology methods and analysis of a nascent Angolan project aimed at preserving the nesting female population of the Olive
[...] Read more.
The scarcity on the Atlantic coast of the African sea turtle population and its dynamics data is well known. This article discusses the nesting ecology methods and analysis of a nascent Angolan project aimed at preserving the nesting female population of the Olive Ridley turtle (Lepidochelys olivacea) on the coast of Lobito. This study examines the nesting ecology of this species from 2020 to 2023. Females had an average CCL of 70.2 cm and CCW of 68.5 cm. These females laid 127 eggs in nests that averaged 47.0 cm deep. The ex situ nest incubation period averaged 60 days, and the hatchling success was 82.1%. Some techniques used in this project require modifications and enhancements. The utilization of photo identification did not yield the anticipated outcomes, prompting the adoption of passive integrated transponders (PITs) in the last season. However, due to limited funding, the success of this method is contingent upon an augmented field effort, allowing for the recapture of a larger number of females. The continuity of this project hinges upon collaboration between higher authorities and the local community. Together, it is possible to deepen the understanding of the nesting ecology of this species and address pivotal issues for its conservation, thereby implementing the most effective preservation measures.
Full article
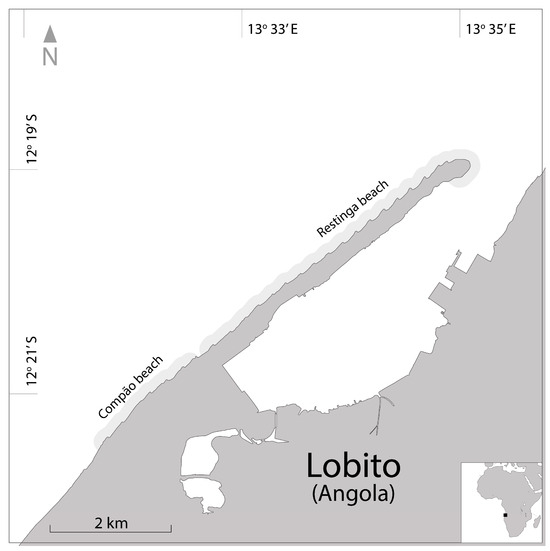
Figure 1
Open AccessCommunication
Valorizing Tree-Nutshell Particles as Delivery Vehicles for a Natural Herbicide
by
, , , , , , and
Methods Protoc. 2024, 7(1), 1; https://doi.org/10.3390/mps7010001 - 20 Dec 2023
Abstract
►▼
Show Figures
The United States is a principal producer of tree nuts (almonds, pistachios, and walnuts), resulting in the generation of excess of tree-nutshell by-products each year, with few market outlets. A nutshell is an essential, lignocellulosic layer that protects a kernel (seed) from the
[...] Read more.
The United States is a principal producer of tree nuts (almonds, pistachios, and walnuts), resulting in the generation of excess of tree-nutshell by-products each year, with few market outlets. A nutshell is an essential, lignocellulosic layer that protects a kernel (seed) from the environment during cultivation. The objective of this study was to develop nutshell by-products as herbicide delivery systems, which would not only enable sustainable weed control in fields but also increases nutshell value and reduce the cost of waste disposal. We recently identified a natural salicylaldehyde (SA) that emits volatiles with both herbicidal and antifungal properties. In this study, walnut shell particles saturated with 0.8 to 1.6 M SA were developed as delivery vehicles for SA to soil, which allowed for the controlled release of an SA fumigant for weed control. The pre- and post-emergent herbicidal efficacy of SA was investigated using model monocot (Lolium arundinaceum (Schreb.) Darbysh; turfgrass) and dicot (Brassica rapa var. pekinensis; Chinese cabbage) plants. We compared (1) the effects of different types of solvents for dissolving SA (dimethyl sulfoxide (DMSO) and ethanol (60%, v/v)), and (2) the effect of covering soil with plastic layers (i.e., soil pasteurization) or not covering soil during SA fumigation using nutshells. Results: In the pre-emergent herbicidal testing with the soil covered, the dicot plants exhibited levels of higher susceptibility to SA in DMSO emitted from nutshells when compared to the monocot plants. The seed germination frequencies in the dicots were 15% and 1% with 0.8 and 1.6 M SA, respectively, while those in the monocots were 32% and 18%, respectively, under the same test conditions. In the post-emergent herbicidal testing with the soil covered, the growth of both the monocot and dicot plants was completely prevented after 5 to 7 days of SA fumigation, resulting in the deaths of entire plants. It was noteworthy that in the post-emergent herbicidal testing, SA dissolved in ethanol (60%, v/v) completely disrupted the growth of the monocot and dicot plants as early as 3 days after SA emission from the nutshells, even without the soil being covered. Tree-nutshell particles could serve as effective SA delivery vehicles with controlled release capabilities for SA. The SA exhibited pre- and post-emergent herbicidal activities against the monocot and dicot plants at most growth stages. SA (0.8 and 1.6 M) dissolved in ethanol (60%, v/v) might exert a synergism for higher herbicidal activity after emission from nutshells. Since tree nuts capture/store a substantial amount of carbon over their life-cycles, the new and sustainable utility of using nutshells not only reduces carbon emissions but also valorizes tree-nut by-products, thus benefitting the tree-nut industry.
Full article
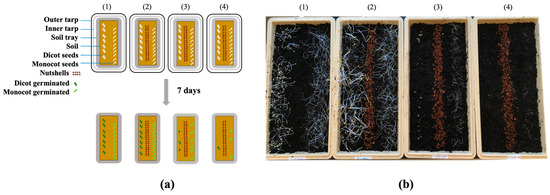

Figure 1
Open AccessTechnical Note
Development of a 3D Perfused In Vitro System to Assess Proangiogenic Properties of Compounds
Methods Protoc. 2023, 6(6), 119; https://doi.org/10.3390/mps6060119 - 09 Dec 2023
Abstract
Perturbation of angiogenesis is associated with a variety of diseases and pro- as well as antiangiogenic therapies are being actively explored. Additionally, unintended adverse drug effects on angiogenesis might lead to promotion of tumor progression and cardiovascular complications. Several tri-dimensional microfluidic vessel-on-chip systems
[...] Read more.
Perturbation of angiogenesis is associated with a variety of diseases and pro- as well as antiangiogenic therapies are being actively explored. Additionally, unintended adverse drug effects on angiogenesis might lead to promotion of tumor progression and cardiovascular complications. Several tri-dimensional microfluidic vessel-on-chip systems have been described that allow a more accurate investigation of vascular physiology and pathology, compared to the two-dimensional static culture of endothelial cells. The OrganoPlate® angiogenesis-on-chip system has been demonstrated to be amenable to high-throughput screening for the antiangiogenic properties of molecules. We set out to adapt this system for high-throughput screening of molecules with proangiogenic properties. Our technical advancement of the OrganoPlate® angiogenesis-on-chip assay expands its applicability in the early screening of both anti- as well as proangiogenic properties of compounds for therapeutic modulation of angiogenesis as well as the identification of angiogenesis-associated drug-induced vascular toxicities.
Full article
(This article belongs to the Section Tissue Engineering and Organoids)
►▼
Show Figures
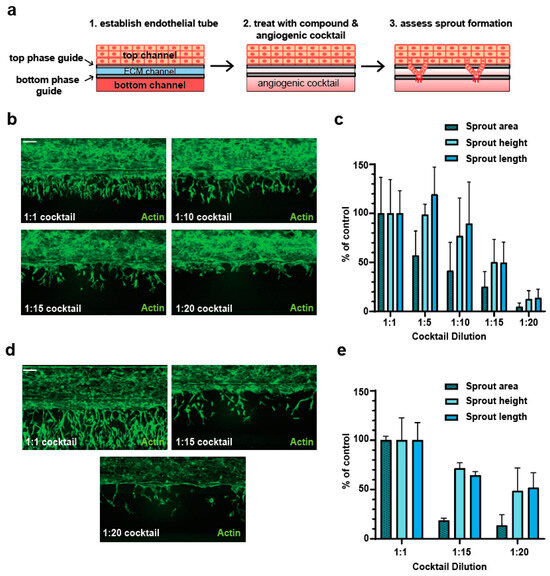
Figure 1
Open AccessCommunication
Magnetic Resonance Imaging as a Tool for Monitoring Intratibial Growth of Experimental Prostate Cancer Metastases in Mice
Methods Protoc. 2023, 6(6), 118; https://doi.org/10.3390/mps6060118 - 05 Dec 2023
Abstract
Bone metastases cause morbidity and mortality in several human cancer forms. Experimental models are used to unravel the mechanisms and identify possible treatment targets. The location inside the skeleton complicates accurate assessment. This study evaluates the performance of magnetic resonance imaging (MRI) of
[...] Read more.
Bone metastases cause morbidity and mortality in several human cancer forms. Experimental models are used to unravel the mechanisms and identify possible treatment targets. The location inside the skeleton complicates accurate assessment. This study evaluates the performance of magnetic resonance imaging (MRI) of prostate cancer tumors growing intratibially in mice. MRI detected intratibial tumor lesions with a sensitivity and specificity of 100% and 89%, respectively, compared to histological evaluation. Location and some phenotypical features could also be readily detected with MRI. Regarding volume estimation, the correlation between MRI and histological assessment was high (p < 0.001, r = 0.936). In conclusion, this study finds MRI to be a reliable tool for in vivo, non-invasive, non-ionizing, real-time monitoring of intratibial tumor growth.
Full article
(This article belongs to the Section Biomedical Sciences and Physiology)
►▼
Show Figures

Figure 1
Open AccessStudy Protocol
The Integrated Health Monitor COVID-19: A Protocol for a Comprehensive Assessment of the Short- and Long-Term Health Impact of the Pandemic in the Netherlands
by
, , , , , , , and
Methods Protoc. 2023, 6(6), 117; https://doi.org/10.3390/mps6060117 - 02 Dec 2023
Abstract
Background: The global COVID-19 pandemic has profoundly affected public health. Directly, the pandemic resulted in over 6.6 million deaths, numerous hospitalizations, and widespread illness. The pandemic has also affected health indirectly through government-imposed protective measures, causing decline in mental well-being and increasing social
[...] Read more.
Background: The global COVID-19 pandemic has profoundly affected public health. Directly, the pandemic resulted in over 6.6 million deaths, numerous hospitalizations, and widespread illness. The pandemic has also affected health indirectly through government-imposed protective measures, causing decline in mental well-being and increasing social isolation. Unlike previous disasters or crises, the pandemic’s worldwide and enduring impact necessitates a unique research approach. The Network for Health Research in Disasters in the Netherlands responded by initiating a longitudinal, extensive research project called the Integrated Health Monitor COVID-19. The Integrated Health Monitor COVID-19 explores both the direct and indirect health effects of the pandemic at the population level. Methods: The Integrated Health Monitor COVID-19 employs a dual-pronged monitoring strategy alongside an annual literature review. This strategy comprises short-cycle monitoring (conducted quarterly) and long-cycle monitoring (conducted once every one or two years). This comprehensive approach enables the evaluation of health trends during the pandemic, facilitating comparisons with pre-pandemic levels and identification of risk and protective factors. Both monitoring methods incorporate data from surveys and general practice registries. The integration of annual literature reviews with these measurements enables iterative research, while dialogues on policy and practice improvements enhance the knowledge-to-action process. Discussion: Much of the existing knowledge about the potential impact of the COVID-19 pandemic is derived from research on sudden-onset disasters limited to specific geographical areas. This study is anticipated to provide valuable fresh insights into the evolving dynamics of population health and specific vulnerabilities within the ongoing pandemic context.
Full article
(This article belongs to the Section Public Health Research)
►▼
Show Figures

Figure 1
Open AccessTechnical Note
Quantitative Image Analysis of Axonal Morphology in In Vivo Model
Methods Protoc. 2023, 6(6), 116; https://doi.org/10.3390/mps6060116 - 01 Dec 2023
Abstract
Quantifying axonal branching is crucial for understanding neural circuit function, developmental and regeneration processes and disease mechanisms. Factors that regulate patterns of axonal arborization and tune neuronal circuits are investigated for their implication in various disorders in brain connectivity. The lack of a
[...] Read more.
Quantifying axonal branching is crucial for understanding neural circuit function, developmental and regeneration processes and disease mechanisms. Factors that regulate patterns of axonal arborization and tune neuronal circuits are investigated for their implication in various disorders in brain connectivity. The lack of a reliable and user-friendly method makes the quantitative analysis of axon morphology difficult. Specifically, methods to visualize and quantify the complex axon arborization are challenging to implement and apply practically. Our study was aimed at developing a robust but simple method of quantification that used ImageJ 2D analysis and compared it with Imaris visualization and analysis of 3D images. We used zebrafish fluorescent transgenic lines to perform in vivo imaging of developing motor neuron axons that adequately reflected the complexity of axonal networks. Our new method, developed on ImageJ, is easy and fast, giving access to new information such as collateral distribution along the axonal shaft. This study describes step-by-step procedures that can be easily applied to a variety of organisms and in vitro systems. Our study provides a basis for further exploration of neural circuits to gain new insights into neuronal disorders and potential therapeutic interventions.
Full article
(This article belongs to the Section Biomedical Sciences and Physiology)
►▼
Show Figures
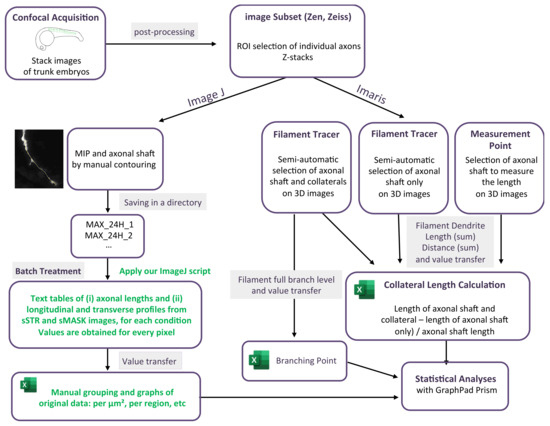
Figure 1
Open AccessProtocol
Protocol Article: A Cross-Sectional Evaluation of Children’s Feet and Lower Extremities
by
, , , , , and
Methods Protoc. 2023, 6(6), 115; https://doi.org/10.3390/mps6060115 - 01 Dec 2023
Abstract
Background: The health of children’s lower extremities and feet is a focus area for caregivers and healthcare professionals such as doctors, school nurses, and podiatrists. Our study aims to investigate the general health status of Danish children’s lower extremities and feet to identify
[...] Read more.
Background: The health of children’s lower extremities and feet is a focus area for caregivers and healthcare professionals such as doctors, school nurses, and podiatrists. Our study aims to investigate the general health status of Danish children’s lower extremities and feet to identify anthropometric parameters that might be preconditions for pain and evaluate for foot diseases and whether they are associated with pain intensity and location, three-dimensional foot dimensions and foot pressure mapping, shoe dimensions, types and intensity of sports activity, quality of life, and foot health. The aim is that we will be able to identify parameters pre-dispositioning for pain, thus providing recommendations for sports activities in relation to the anthropometric conditions of a child as a potential preventive measure for pain. This analysis will be stratified by socioeconomic status on a group level, and this perspective will be able to provide preventative recommendations to prevent pain. Methods: This study is a cross-sectional examination of a thousand children in the first, fifth, and ninth grades in randomized selected Danish primary schools. We will perform a clinical examination of the lower extremities and feet for misalignments, deformities, and diseases as well as rotational status and range of motion. Moreover, we will evaluate their pain levels, sports activities, three-dimensional foot dimensions, plantar pressure, footwear, and patient-related outcome measures (PROMs) for foot health and quality of life. Results: We aim to provide an anthropometrical overview of the lower extremities and feet in children. The obtained basic understanding of healthy normal material in children will be analyzed for its relationships with pain level, sports activities, and socioeconomic status on a group level. This could potentially provide us with an understanding of the factors that impact lower extremity and foot diseases in children. In conclusion, examining children’s lower extremities and feet in Danish primary schools is a step toward identifying areas of improvement in self-care and shoe fitting, mapping podiatry-related needs of care in children’s feet, and providing parental recommendations for preventive actions on shoe fitting and the choice and intensity of sports activity concerning pain. Conclusions: The tenet of this study is a long-term follow-up to evaluate the long-term socioeconomic course on a group level, foot status, and sports activity, using patient-related outcome measures evaluating quality of life and other lifestyle factors such as emotional functioning, social functioning and interaction, and school functioning. Potentially, this will improve children’s quality of life and prevent future diseases.
Full article
(This article belongs to the Section Public Health Research)
►▼
Show Figures
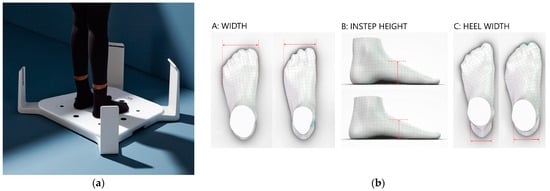
Figure 1
Open AccessArticle
Validity and Reliability of POM-Checker for Measuring Shoulder Range of Motion in Healthy Participants: A Pilot Single-Center Comparative Study
by
, , , , , and
Methods Protoc. 2023, 6(6), 114; https://doi.org/10.3390/mps6060114 - 27 Nov 2023
Abstract
Background. The aim of this study was to compare shoulder movement measurements between a Kinect-based markerless ROM assessment device (POM-Checker) and a 3D motion capture analysis system (BTS SMART DX-400). Methods. This was a single-visit clinical trial designed to evaluate the validity and
[...] Read more.
Background. The aim of this study was to compare shoulder movement measurements between a Kinect-based markerless ROM assessment device (POM-Checker) and a 3D motion capture analysis system (BTS SMART DX-400). Methods. This was a single-visit clinical trial designed to evaluate the validity and reliability of the POM-Checker. The primary outcome was to assess the equivalence between two measurement devices within the same set of participants, aiming to evaluate the validity of the POM-Checker compared to the gold standard device (3D Motion Analysis System). As this was a pilot study, six participants were included. Results. The intraclass correlation coefficient (ICC) and the corresponding 95% confidence intervals (CIs) were used to assess the reproducibility of the measurements. Among the 18 movements analyzed, 16 exhibited ICC values of >0.75, indicating excellent reproducibility. Conclusion. The results showed that the POM-checker is reliable and validated to measure the range of motion of the shoulder joint.
Full article
(This article belongs to the Section Biomedical Sciences and Physiology)
►▼
Show Figures
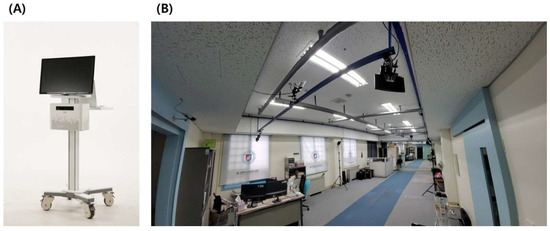

Figure 1
Open AccessProtocol
A Versatile Protocol for Efficient Transformation and Regeneration in Mega Indica Rice Cultivar MTU1010: Optimization through Hormonal Variables
by
, , , , , and
Methods Protoc. 2023, 6(6), 113; https://doi.org/10.3390/mps6060113 - 23 Nov 2023
Abstract
Rice is one of the apex food crops in terms of meeting the daily calorific and dietary requirement of the majority of the world population. However, rice productivity is severely limited by various biotic and abiotic attributes, causing a severe threat to global
[...] Read more.
Rice is one of the apex food crops in terms of meeting the daily calorific and dietary requirement of the majority of the world population. However, rice productivity is severely limited by various biotic and abiotic attributes, causing a severe threat to global food security. In the use of functional genomics and genome editing for the generation of trait-enhanced genotypes, it is necessary to have an efficient genetic transformation and regeneration protocol. The recalcitrant nature and paucity of efficient and versatile genetic transformation and regeneration protocols for indica cultivars remains a constraint. In the present study, we have optimized a tissue culture method for MTU1010, a mega indica rice variety. We conducted a combinatorial analysis of different plant growth regulators on embryogenic callus induction efficiency, and it was observed that MSB5 medium supplemented with 2.5 mg/L 2-4D and 0.25 mg/L 6-BAP results in maximum embryogenic callus induction, i.e., 92%. The regeneration efficiency of a transformed callus can be enhanced by up to 50% with the supplementation of 1 mg/L kinetin alongside 2.5 mg/L BAP and 0.5 mg/L NAA in the shooting medium. Furthermore, our results unveiled that the pre-activation of Agrobacterium culture for 30 min with 150 µM acetosyringone significantly increased the transformation efficiency of calli. Additionally, descaling the salt concentration to half strength in resuspension and co-cultivation increased the efficiency of transformation up to 33%. Thus, the protocol developed in this study will be instrumental for the genome editing and genetic engineering of indica rice cultivars for functional genomics studies and crop improvement.
Full article
(This article belongs to the Collection Gene Editing)
►▼
Show Figures
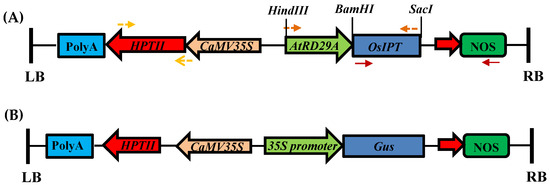
Figure 1
Open AccessProtocol
A Simple, Rapid, and Effective Heparinase Protocol to Enable Nucleic Acid Study from Frozen Heparinized Plasma
by
, , , , , , , , and
Methods Protoc. 2023, 6(6), 112; https://doi.org/10.3390/mps6060112 - 20 Nov 2023
Abstract
►▼
Show Figures
Cell-free RNAs (cfRNAs) are promising analytes as non-invasive biomarkers and have even greater potential if tied in with metabolomics. Plasma is an optimal source for cfRNAs but is often derived from a variety of anticoagulants. Plasma obtained in heparin is suitable for metabolomics
[...] Read more.
Cell-free RNAs (cfRNAs) are promising analytes as non-invasive biomarkers and have even greater potential if tied in with metabolomics. Plasma is an optimal source for cfRNAs but is often derived from a variety of anticoagulants. Plasma obtained in heparin is suitable for metabolomics but is difficult to utilize for qPCR-based downstream analysis. In the present study, we aimed to develop a simple, time-efficient, and cost-effective heparinase protocol, followed by library preparation and sequencing of human plasma cfRNAs drawn and stored in heparin at −80 °C for several years. Blood was collected in CPT™ sodium heparin tubes from patients with chronic HCV infection (NCT02400216) at the National Institutes of Health (NIH) Clinical Center. Plasma cfRNAs were treated with heparinase I and used for library preparation and next-generation sequencing (NGS). Heparinase treatment maintained RNA integrity and allowed for successful library preparation for all the study subjects even with 7 ng of cfRNAs as starting material. The classification report derived from Pavian R package v1.2.0 showed no artificial reads. The abundance of chordate over microbial reads suggests no addition of experimental error through heparinase I treatment. We report a novel and practical approach to heparinase treatment for human plasma collected and frozen in sodium heparin for several years. This is an effective demonstration of utilizing heparin plasma for NGS and downstream transcriptomic research, which could then be integrated with metabolomics from the same samples, maximizing efficiency and minimizing blood draws.
Full article

Figure 1
Open AccessEditorial
Methods and Protocols—Aims and Scope Update
by
and
Methods Protoc. 2023, 6(6), 111; https://doi.org/10.3390/mps6060111 - 17 Nov 2023
Abstract
As our readers know, Methods and Protocols is a multidisciplinary peer-reviewed scientific journal that provides a forum to the publication of novel approaches in the fields of Life Sciences, Chemistry, and Biomedical Sciences and their intersection with other related scientific fields such as
[...] Read more.
As our readers know, Methods and Protocols is a multidisciplinary peer-reviewed scientific journal that provides a forum to the publication of novel approaches in the fields of Life Sciences, Chemistry, and Biomedical Sciences and their intersection with other related scientific fields such as Physics, Earth Sciences, and Environmental Research [...]
Full article
Open AccessProtocol
Protocol for Facile Synthesis of Fmoc-N-Me-AA-OH Using 2-CTC Resin as Temporary and Reusable Protecting Group
by
, , , , and
Methods Protoc. 2023, 6(6), 110; https://doi.org/10.3390/mps6060110 - 13 Nov 2023
Abstract
One approach to enhance the bioavailability and half-life of peptides in vivo is through N-methylation of one or more of the amino acids within the peptide sequence. However, commercially available Fmoc-N-Me-AA-OHs are limited and often expensive. In this study, a solid-phase synthesis method
[...] Read more.
One approach to enhance the bioavailability and half-life of peptides in vivo is through N-methylation of one or more of the amino acids within the peptide sequence. However, commercially available Fmoc-N-Me-AA-OHs are limited and often expensive. In this study, a solid-phase synthesis method for Fmoc-N-Me-AA-OH was developed using a 2-chlorotrityl chloride (2-CTC) resin as a temporary protective group for the carboxylic acid strategy. Two strategies for the alkylation step were compared, employing either dimethyl sulfate or methyl iodide in the Biron−Kessler method. In this work we tested the protocol with two amino acids: Fmoc-Thr(tBu)-OH and Fmoc-βAla-OH. The first one is an alpha amino acid, very hindered and with the amine group directly influenced by the electronic effects of the carboxy group, whereas in Fmoc-βAla-OH, the presence of a methylene group weakens this influence due to the intervening carbon atoms. The desired amino acids, Fmoc-N-Me-Thr(tBu)-OH and Fmoc-N-Me-βAla-OH, were synthesized by both strategies with high yield and purity.
Full article
(This article belongs to the Special Issue Feature Papers in Methods and Protocols 2023)
►▼
Show Figures
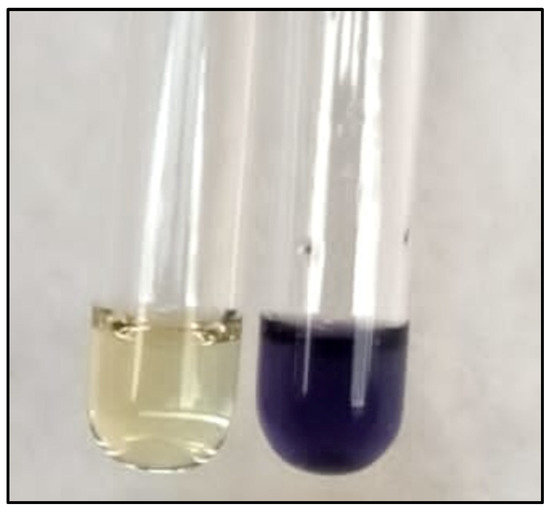

Figure 1
Open AccessProtocol
Bio-SELEX: A Strategy for Biomarkers Isolation Directly from Biological Samples
Methods Protoc. 2023, 6(6), 109; https://doi.org/10.3390/mps6060109 - 11 Nov 2023
Abstract
►▼
Show Figures
Bio-SELEX is a revolutionary method for the discovery of novel biomarkers within biological samples, offering profound insights into diagnosing both infectious and non-infectious diseases. This innovative strategy involves three crucial steps: Traditional SELEX, Pull Down, and mass spectrometry. Firstly, Traditional SELEX involves the
[...] Read more.
Bio-SELEX is a revolutionary method for the discovery of novel biomarkers within biological samples, offering profound insights into diagnosing both infectious and non-infectious diseases. This innovative strategy involves three crucial steps: Traditional SELEX, Pull Down, and mass spectrometry. Firstly, Traditional SELEX involves the systematic selection of specific nucleic acid sequences (aptamers) that bind to the target molecules of interest. These aptamers are generated through iterative rounds of selection, amplification, and enrichment, ultimately yielding highly selective ligands. Secondly, the Pull-Down phase employs these aptamers to capture and isolate the target biomarkers from complex biological samples. This step ensures the specificity of the selected aptamers in binding to their intended targets. Lastly, mass spectrometry is utilized to identify and quantify the captured biomarkers, providing precise information about their presence and concentration in the sample. These quantitative data are invaluable in disease diagnosis and monitoring. Bio-SELEX’s significance lies in its ability to discover biomarkers for a wide range of diseases, spanning infectious and non-infectious conditions. This approach holds great promise for early disease detection, personalized medicine, and the development of targeted therapies. By harnessing the power of aptamers and mass spectrometry, Bio-SELEX advances our understanding of disease biology and opens new avenues for improved healthcare.
Full article

Figure 1
Open AccessBrief Report
Comparison of Light-Sheet Fluorescence Microscopy and Fast-Confocal Microscopy for Three-Dimensional Imaging of Cleared Mouse Brain
Methods Protoc. 2023, 6(6), 108; https://doi.org/10.3390/mps6060108 - 10 Nov 2023
Abstract
►▼
Show Figures
Whole-brain imaging is important for understanding brain functions through deciphering tissue structures, neuronal circuits, and single-neuron tracing. Thus, many clearing methods have been developed to acquire whole-brain images or images of three-dimensional thick tissues. However, there are several limitations to imaging whole-brain volumes,
[...] Read more.
Whole-brain imaging is important for understanding brain functions through deciphering tissue structures, neuronal circuits, and single-neuron tracing. Thus, many clearing methods have been developed to acquire whole-brain images or images of three-dimensional thick tissues. However, there are several limitations to imaging whole-brain volumes, including long image acquisition times, large volumes of data, and a long post-image process. Based on these limitations, many researchers are unsure about which light microscopy is most suitable for imaging thick tissues. Here, we compared fast-confocal microscopy with light-sheet fluorescence microscopy for whole-brain three-dimensional imaging, which can acquire images the fastest. To compare the two types of microscopies for large-volume imaging, we performed tissue clearing of a whole mouse brain, and changed the sample chamber and low- magnification objective lens and modified the sample holder of a light-sheet fluorescence microscope. We found out that light-sheet fluorescence microscopy using a 2.5× objective lens possesses several advantages, including saving time, large-volume image acquisitions, and high Z-resolution, over fast-confocal microscopy, which uses a 4× objective lens. Therefore, we suggest that light-sheet fluorescence microscopy is suitable for whole mouse brain imaging and for obtaining high-resolution three-dimensional images.
Full article

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Sensors, Viruses, Diseases, Algorithms, MPs, Symmetry
Computational Intelligence for Virus and Bacteria Detection in Multi Surface Environments
Topic Editors: Marcin Woźniak, Muhammad Fazal IjazDeadline: 31 January 2024

Conferences
Special Issues
Special Issue in
MPs
Biosensors Based on Recombinant Antibodies—in Memory of Professor Hiroshi UedaGuest Editors: Peter Kristensen, Jinhua DongDeadline: 20 May 2024
Special Issue in
MPs
Plant Tissue Culture for Crop Improvement
Guest Editor: Pankaj BhowmikDeadline: 20 August 2024
Special Issue in
MPs
Methods on Sport Biomechanics
Guest Editors: Didier Pradon, Jean Slawinski, Fabien LeboeufDeadline: 20 December 2024
Special Issue in
MPs
Feature Papers in Methods and Protocols 2024
Guest Editor: Fernando AlbericioDeadline: 31 December 2024
Topical Collections
Topical Collection in
MPs
Green Chemistry
Collection Editors: Artur M. S. Silva, Diana Cláudia Pinto
Topical Collection in
MPs
Indigenous Health
Collection Editors: Juergen Reichardt, Aunty Kerrie Doyle













