Journal Description
Non-Coding RNA
Non-Coding RNA
is an international, peer-reviewed, open access journal on non-coding RNA research dealing with elucidating the structure, function and biology of regulatory non-coding RNAs. Non-Coding RNA is published bimonthly online by MDPI. Cardiolinc is affiliated with Non-Coding RNA.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, CAPlus / SciFinder, and other databases.
- Journal Rank: CiteScore - Q1 (Genetics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.6 days after submission; acceptance to publication is undertaken in 3.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
4.3 (2022)
Latest Articles
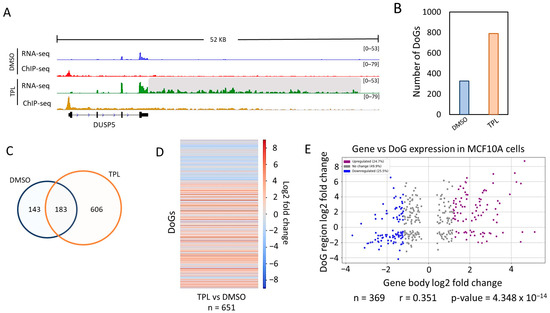
Transcriptional Stress Induces the Generation of DoGs in Cancer Cells
Non-Coding RNA 2024, 10(1), 5; https://doi.org/10.3390/ncrna10010005 - 10 Jan 2024
Abstract
►
Show Figures
A characteristic of the cellular response to stress is the production of RNAs generated from a readthrough transcription of genes, called downstream-of-gene-(DoG)-containing transcripts. Additionally, transcription inhibitor drugs are candidates for fighting cancer. In this work, we report the results of a bioinformatic analysis
[...] Read more.
A characteristic of the cellular response to stress is the production of RNAs generated from a readthrough transcription of genes, called downstream-of-gene-(DoG)-containing transcripts. Additionally, transcription inhibitor drugs are candidates for fighting cancer. In this work, we report the results of a bioinformatic analysis showing that one of the responses to transcription inhibition is the generation of DoGs in cancer cells. Although some genes that form DoGs were shared between the two cancer lines, there did not appear to be a functional correlation between them. However, our findings show that DoGs are generated as part of the cellular response to transcription inhibition like other types of cellular stress, suggesting that they may be part of the defense against transcriptional stress.
Full article
Open AccessCommunication
Genetic Loss of miR-205 Causes Increased Mammary Gland Development
by
, , , , , , and
Non-Coding RNA 2024, 10(1), 4; https://doi.org/10.3390/ncrna10010004 - 31 Dec 2023
Abstract
MiRNAs play crucial roles in a broad spectrum of biological processes, both physiological and pathological. Different reports implicate miR-205 in the control of breast stem cell properties. Differential miR-205 expression has been observed in different stages of mammary gland development and maturation. However,
[...] Read more.
MiRNAs play crucial roles in a broad spectrum of biological processes, both physiological and pathological. Different reports implicate miR-205 in the control of breast stem cell properties. Differential miR-205 expression has been observed in different stages of mammary gland development and maturation. However, a functional role in this process has not been clearly demonstrated. We generated an miR-205 knockout in the FVB/N mouse strain, which is viable and characterized by enhanced mammary gland development. Indeed, mammary glands of miR-205−/− female mice at different ages (1.5 and 5.5 months) show increased outgrowth and branching. This evidence is consistent with our previously reported data demonstrating the direct miR-205-mediated targeting of HER3, a master regulator of mammary gland development, and the oncosuppressive activity of this microRNA in different types of breast cancer.
Full article
(This article belongs to the Topic MicroRNA: Mechanisms of Action, Physio-Pathological Implications, and Disease Biomarkers, 2nd Volume)
►▼
Show Figures
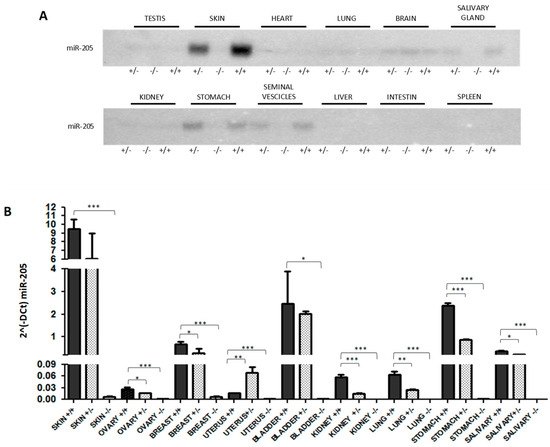
Figure 1
Open AccessReview
Long Non-Coding RNAs (lncRNAs) in Heart Failure: A Comprehensive Review
by
, , , and
Non-Coding RNA 2024, 10(1), 3; https://doi.org/10.3390/ncrna10010003 - 28 Dec 2023
Abstract
Heart failure (HF) is a widespread cardiovascular condition that poses significant risks to a wide spectrum of age groups and leads to terminal illness. Although our understanding of the underlying mechanisms of HF has improved, the available treatments still remain inadequate. Recently, long
[...] Read more.
Heart failure (HF) is a widespread cardiovascular condition that poses significant risks to a wide spectrum of age groups and leads to terminal illness. Although our understanding of the underlying mechanisms of HF has improved, the available treatments still remain inadequate. Recently, long non-coding RNAs (lncRNAs) have emerged as crucial players in cardiac function, showing possibilities as potential targets for HF therapy. These versatile molecules interact with chromatin, proteins, RNA, and DNA, influencing gene regulation. Notable lncRNAs like Fendrr, Trpm3, and Scarb2 have demonstrated therapeutic potential in HF cases. Additionally, utilizing lncRNAs to forecast survival rates in HF patients and distinguish various cardiac remodeling conditions holds great promise, offering significant benefits in managing cardiovascular disease and addressing its far-reaching societal and economic impacts. This underscores the pivotal role of lncRNAs in the context of HF research and treatment.
Full article
(This article belongs to the Special Issue Non-coding RNAs: Multiple Players in Human Diseases)
►▼
Show Figures
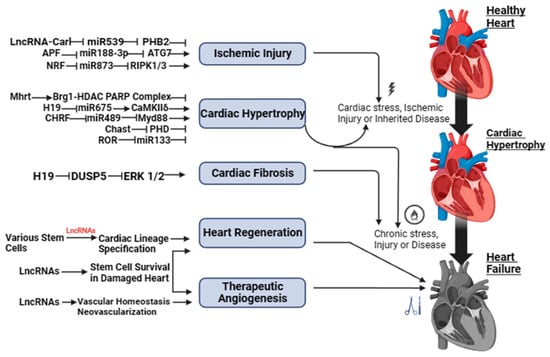
Figure 1
Open AccessArticle
MiR-4646-5p Acts as a Tumor-Suppressive Factor in Triple Negative Breast Cancer and Targets the Cholesterol Transport Protein GRAMD1B
by
, , , , , , , , and
Non-Coding RNA 2024, 10(1), 2; https://doi.org/10.3390/ncrna10010002 - 26 Dec 2023
Abstract
MicroRNAs (miRNAs) are crucial post-transcriptional regulators of gene expression, and their deregulation contributes to many aspects of cancer development and progression. Thus, miRNAs provide insight into oncogenic mechanisms and represent promising targets for new therapeutic approaches. A type of cancer that is still
[...] Read more.
MicroRNAs (miRNAs) are crucial post-transcriptional regulators of gene expression, and their deregulation contributes to many aspects of cancer development and progression. Thus, miRNAs provide insight into oncogenic mechanisms and represent promising targets for new therapeutic approaches. A type of cancer that is still in urgent need of improved treatment options is triple negative breast cancer (TNBC). Therefore, we aimed to characterize a novel miRNA with a potential role in TNBC. Based on a previous study, we selected miR-4646-5p, a miRNA with a still unknown function in breast cancer. We discovered that higher expression of miR-4646-5p in TNBC patients is associated with better survival. In vitro assays showed that miR-4646-5p overexpression reduces growth, proliferation, and migration of TNBC cell lines, whereas inhibition had the opposite effect. Furthermore, we found that miR-4646-5p inhibits the tube formation ability of endothelial cells, which may indicate anti-angiogenic properties. By whole transcriptome analysis, we not only observed that miR-4646-5p downregulates many oncogenic factors, like tumor-promoting cytokines and migration- and invasion-related genes, but were also able to identify a direct target, the GRAM domain-containing protein 1B (GRAMD1B). GRAMD1B is involved in cellular cholesterol transport and its knockdown phenocopied the growth-reducing effects of miR-4646-5p. We thus conclude that GRAMD1B may partly contribute to the diverse tumor-suppressive effects of miR-4646-5p in TNBC.
Full article
(This article belongs to the Special Issue Molecular Mechanisms and Clinical Implications of Non-coding RNAs in Cancer)
►▼
Show Figures
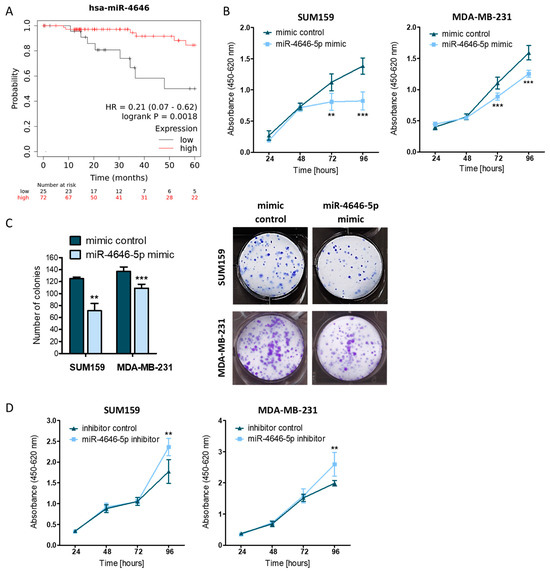
Figure 1
Open AccessArticle
Sequencing Reveals miRNAs Enriched in the Developing Mouse Enteric Nervous System
Non-Coding RNA 2024, 10(1), 1; https://doi.org/10.3390/ncrna10010001 - 22 Dec 2023
Abstract
The enteric nervous system (ENS) is an essential network of neurons and glia in the bowel wall. Defects in ENS development can result in Hirschsprung disease (HSCR), a life-threatening condition characterized by severe constipation, abdominal distention, bilious vomiting, and failure to thrive. A
[...] Read more.
The enteric nervous system (ENS) is an essential network of neurons and glia in the bowel wall. Defects in ENS development can result in Hirschsprung disease (HSCR), a life-threatening condition characterized by severe constipation, abdominal distention, bilious vomiting, and failure to thrive. A growing body of literature connects HSCR to alterations in miRNA expression, but there are limited data on the normal miRNA landscape in the developing ENS. We sequenced small RNAs (smRNA-seq) and messenger RNAs (mRNA-seq) from ENS precursor cells of mid-gestation Ednrb-EGFP mice and compared them to aggregated RNA from all other cells in the developing bowel. Our smRNA-seq results identified 73 miRNAs that were significantly enriched and highly expressed in the developing ENS, with miR-9, miR-27b, miR-124, miR-137, and miR-488 as our top 5 miRNAs that are conserved in humans. However, contrary to prior reports, our follow-up analyses of miR-137 showed that loss of Mir137 in Nestin-cre, Wnt1-cre, Sox10-cre, or Baf53b-cre lineage cells had no effect on mouse survival or ENS development. Our data provide important context for future studies of miRNAs in HSCR and other ENS diseases and highlight open questions about facility-specific factors in development.
Full article
(This article belongs to the Special Issue Non-coding RNA in the USA: Latest Advances and Perspectives)
►▼
Show Figures
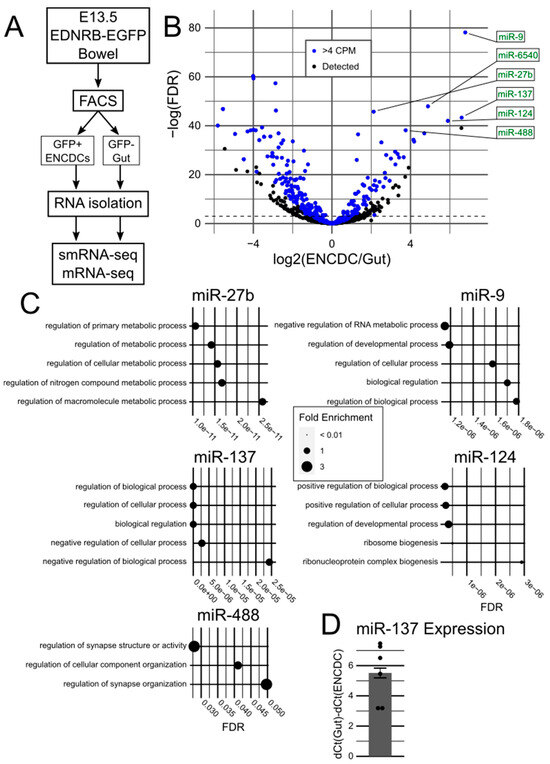
Figure 1
Open AccessEditorial
The Non-Coding RNA Journal Club: Highlights on Recent Papers—13
by
, , , , , , , , , , and
Non-Coding RNA 2023, 9(6), 76; https://doi.org/10.3390/ncrna9060076 - 14 Dec 2023
Abstract
We are delighted to share with you our thirteenth Journal Club and highlight some of the most interesting papers published recently [...]
Full article
(This article belongs to the Collection The Non-Coding RNA Journal Club: Highlights on Recent Papers)
Open AccessFeature PaperReview
Regulation of Macrophage Polarization in Allergy by Noncoding RNAs
Non-Coding RNA 2023, 9(6), 75; https://doi.org/10.3390/ncrna9060075 - 11 Dec 2023
Abstract
Allergy is a type 2 immune reaction triggered by antigens known as allergens, including food and environmental substances such as peanuts, plant pollen, fungal spores, and the feces and debris of mites and insects. Macrophages are myeloid immune cells with phagocytic abilities that
[...] Read more.
Allergy is a type 2 immune reaction triggered by antigens known as allergens, including food and environmental substances such as peanuts, plant pollen, fungal spores, and the feces and debris of mites and insects. Macrophages are myeloid immune cells with phagocytic abilities that process exogenous and endogenous antigens. Upon activation, they can produce effector molecules such as cytokines as well as anti-inflammatory molecules. The dysregulation of macrophage function can lead to excessive type 1 inflammation as well as type 2 inflammation, which includes allergic reactions. Thus, it is important to better understand how macrophages are regulated in the pathogenesis of allergies. Emerging evidence highlights the role of noncoding RNAs (ncRNAs) in macrophage polarization, which in turn can modify the pathogenesis of various immune-mediated diseases, including allergies. This review summarizes the current knowledge regarding this topic and considers three classes of ncRNAs: microRNAs, long ncRNAs, and circular ncRNAs. Understanding the roles of these ncRNAs in macrophage polarization will provide new insights into the pathogenesis of allergies and identify potential novel therapeutic targets.
Full article
(This article belongs to the Special Issue Non-Coding RNA in the Immune System)
►▼
Show Figures
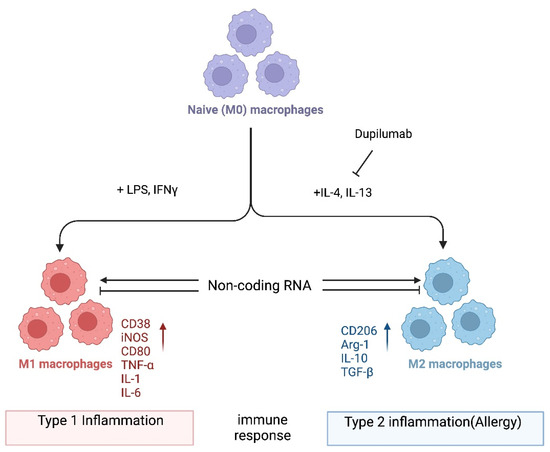
Figure 1
Open AccessArticle
Age and 17β-Estradiol (E2) Facilitate Nuclear Export and Argonaute Loading of microRNAs in the Female Brain
Non-Coding RNA 2023, 9(6), 74; https://doi.org/10.3390/ncrna9060074 - 06 Dec 2023
Abstract
Aging in women is accompanied by a dramatic change in circulating sex steroid hormones. Specifically, the primary circulating estrogen, 17β-estradiol (E2), is nearly undetectable in post-menopausal women. This decline is associated with a variety of cognitive and mood disorders, yet hormone
[...] Read more.
Aging in women is accompanied by a dramatic change in circulating sex steroid hormones. Specifically, the primary circulating estrogen, 17β-estradiol (E2), is nearly undetectable in post-menopausal women. This decline is associated with a variety of cognitive and mood disorders, yet hormone replacement therapy is only effective within a narrow window of time surrounding the menopausal transition. Our previous work identified microRNAs as a potential molecular substrate underlying the change in E2 efficacy associated with menopause in advanced age. Specifically, we showed that E2 regulated a small subset of mature miRNAs in the aging female brain. In this study, we hypothesized that E2 regulates the stability of mature miRNAs by altering their subcellular localization and their association with argonaute proteins. We also tested the hypothesis that the RNA binding protein, hnRNP A1, was an important regulator of mature miR-9-5p expression in neuronal cells. Our results demonstrated that E2 treatment affected miRNA subcellular localization and its association with argonaute proteins differently, depending on the length of time following E2 deprivation (i.e., ovariectomy). We also provide strong evidence that hnRNP A1 regulates the transcription of pri-miR-9 and likely plays a posttranscriptional role in mature miR-9-5p turnover. Taken together, these data have important implications for considering the optimal timing for hormone replacement therapy, which might be less dependent on age and more related to how long treatment is delayed following menopause.
Full article
(This article belongs to the Special Issue Non-coding RNA in the USA: Latest Advances and Perspectives)
►▼
Show Figures
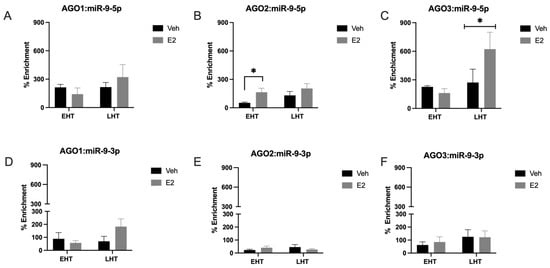
Figure 1
Open AccessArticle
MicroRNAs in Gingival Crevicular Fluid: An Observational Case-Control Study of Differential Expression in Periodontitis
by
, , , , and
Non-Coding RNA 2023, 9(6), 73; https://doi.org/10.3390/ncrna9060073 - 18 Nov 2023
Abstract
►▼
Show Figures
Objectives: microRNAs (miRNAs) present in the gingival crevicular fluid (GCF) of patients with chronic periodontitis may serve as biomarkers of periodontal disease. The aim of this study was to perform a miRNA-sequencing study of all miRNAs present in GCF, comparing miRNA expression level
[...] Read more.
Objectives: microRNAs (miRNAs) present in the gingival crevicular fluid (GCF) of patients with chronic periodontitis may serve as biomarkers of periodontal disease. The aim of this study was to perform a miRNA-sequencing study of all miRNAs present in GCF, comparing miRNA expression level profiles between advanced chronic periodontitis (CP) patients and healthy subjects (HS). Materials and methods: GCF samples were collected from the single-rooted teeth of patients with severe CP (n = 11) and of HS (n = 12). miRNAs were isolated from GCF using an miRNeasy Serum/Plasma kit(Qiagen GmbH, Hilden, Germany). Reverse transcription polymerase chain reaction (qRT-PCR) was used to determine the expression levels of miRNA candidates involved in periodontal pathogenesis. Results: Of all the sequenced miRNAs, miR-199, miR-146a, miR-30a, and miR-338 were identified as best representing the CP patient samples. The validation study identified miR-199 as the most powerful biomarker used to define periodontitis. Conclusions: Upon sequencing all known miRNAs in GCF for the first time, we uncovered several potential biomarkers to define periodontitis. Identifying miRNAS in the GCF using high-throughput approaches will clarify the role of these molecules in periodontitis and provide biomarkers with potential applications.
Full article
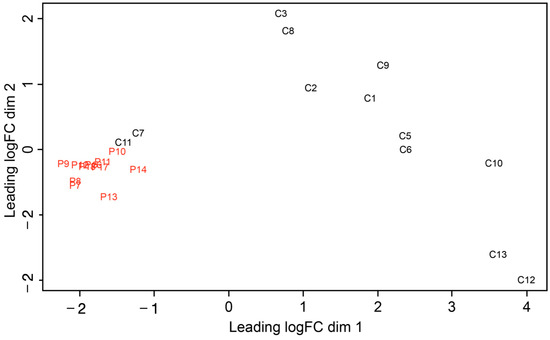
Figure 1
Open AccessArticle
Ethanol- and PARP-Mediated Regulation of Ribosome-Associated Long Non-Coding RNA (lncRNA) in Pyramidal Neurons
Non-Coding RNA 2023, 9(6), 72; https://doi.org/10.3390/ncrna9060072 - 17 Nov 2023
Abstract
►▼
Show Figures
Although, by definition, long noncoding RNAs (lncRNAs) are not translated, they are sometimes associated with ribosomes. In fact, some estimates suggest the existence of more than 50 K lncRNA molecules that could encode for small peptides. We examined the effects of an ethanol
[...] Read more.
Although, by definition, long noncoding RNAs (lncRNAs) are not translated, they are sometimes associated with ribosomes. In fact, some estimates suggest the existence of more than 50 K lncRNA molecules that could encode for small peptides. We examined the effects of an ethanol and Poly-ADP Ribose Polymerase (PARP) inhibitor (ABT-888) on ribosome-bound lncRNAs. Mice were administered via intraperitoneal injection (i.p.) either normal saline (CTL) or ethanol (EtOH) twice a day for four consecutive days. On the fourth day, a sub-group of mice administered with ethanol also received ABT-888 (EtOH+ABT). Ribosome-bound lncRNAs in CaMKIIα-expressing pyramidal neurons were measured using the Translating Ribosome Affinity Purification (TRAP) technique. Our findings show that EtOH altered the attachment of 107 lncRNA transcripts, while EtOH+ABT altered 60 lncRNAs. Among these 60 lncRNAs, 49 were altered by both conditions, while EtOH+ABT uniquely altered the attachment of 11 lncRNA transcripts that EtOH alone did not affect. To validate these results, we selected eight lncRNAs (Mir124-2hg, 5430416N02Rik, Snhg17, Snhg12, Snhg1, Mir9-3hg, Gas5, and 1110038B12Rik) for qRT-PCR analysis. The current study demonstrates that ethanol-induced changes in lncRNA attachment to ribosomes can be mitigated by the addition of the PARP inhibitor ABT-888.
Full article
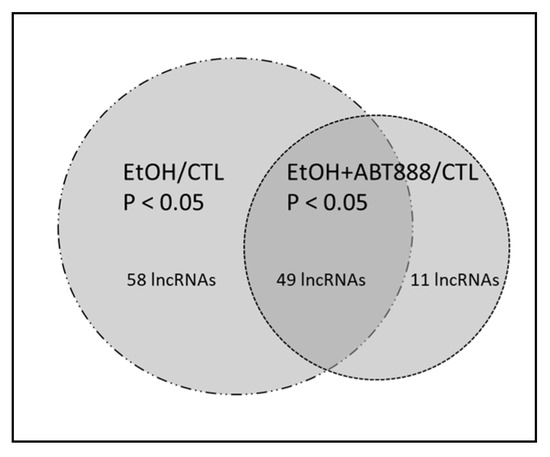
Figure 1
Open AccessArticle
LADON, a Natural Antisense Transcript of NODAL, Promotes Tumour Progression and Metastasis in Melanoma
by
, , , , , and
Non-Coding RNA 2023, 9(6), 71; https://doi.org/10.3390/ncrna9060071 - 15 Nov 2023
Abstract
The TGFβ family member NODAL, repeatedly required during embryonic development, has also been associated with tumour progression. Our aim was to clarify the controversy surrounding its involvement in melanoma tumour progression. We found that the deletion of the NODAL exon 2 in a
[...] Read more.
The TGFβ family member NODAL, repeatedly required during embryonic development, has also been associated with tumour progression. Our aim was to clarify the controversy surrounding its involvement in melanoma tumour progression. We found that the deletion of the NODAL exon 2 in a metastatic melanoma cell line impairs its ability to form tumours and colonize distant tissues. However, we show that this phenotype does not result from the absence of NODAL, but from a defect in the expression of a natural antisense transcript of NODAL, here called LADON. We show that LADON expression is specifically activated in metastatic melanoma cell lines, that its transcript is packaged in exosomes secreted by melanoma cells, and that, via its differential impact on the expression of oncogenes and tumour suppressors, it promotes the mesenchymal to amoeboid transition that is critical for melanoma cell invasiveness. LADON is, therefore, a new player in the regulatory network governing tumour progression in melanoma and possibly in other types of cancer.
Full article
(This article belongs to the Special Issue The Importance of Non-coding RNAs in Epithelial Cancers)
►▼
Show Figures

Figure 1
Open AccessReview
An Overview of the Immune Modulatory Properties of Long Non-Coding RNAs and Their Potential Use as Therapeutic Targets in Cancer
by
, , , and
Non-Coding RNA 2023, 9(6), 70; https://doi.org/10.3390/ncrna9060070 - 11 Nov 2023
Abstract
Long non-coding RNAs (lncRNAs) play pivotal roles in regulating immune responses, immune cell differentiation, activation, and inflammatory processes. In cancer, they are gaining prominence as potential therapeutic targets due to their ability to regulate immune checkpoint molecules and immune-related factors, suggesting avenues for
[...] Read more.
Long non-coding RNAs (lncRNAs) play pivotal roles in regulating immune responses, immune cell differentiation, activation, and inflammatory processes. In cancer, they are gaining prominence as potential therapeutic targets due to their ability to regulate immune checkpoint molecules and immune-related factors, suggesting avenues for bolstering anti-tumor immune responses. Here, we explore the mechanistic insights into lncRNA-mediated immune modulation, highlighting their impact on immunity. Additionally, we discuss their potential to enhance cancer immunotherapy, augmenting the effectiveness of immune checkpoint inhibitors and adoptive T cell therapies. LncRNAs as therapeutic targets hold the promise of revolutionizing cancer treatments, inspiring further research in this field with substantial clinical implications.
Full article
(This article belongs to the Special Issue ncRNAs to Target Molecular Pathways)
►▼
Show Figures
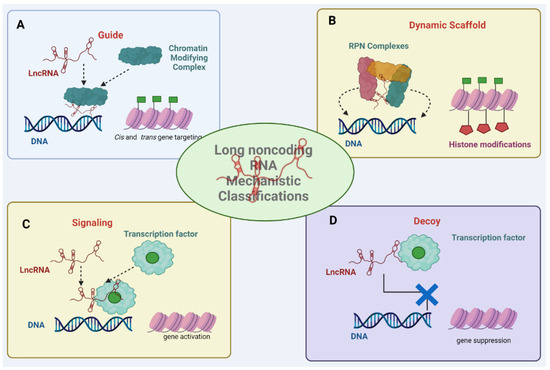
Figure 1
Open AccessArticle
The Typical tRNA Co-Expresses Multiple 5′ tRNA Halves Whose Sequences and Abundances Depend on Isodecoder and Isoacceptor and Change with Tissue Type, Cell Type, and Disease
by
, , , , and
Non-Coding RNA 2023, 9(6), 69; https://doi.org/10.3390/ncrna9060069 - 06 Nov 2023
Abstract
Transfer RNA-derived fragments (tRFs) are noncoding RNAs that arise from either mature transfer RNAs (tRNAs) or their precursors. One important category of tRFs comprises the tRNA halves, which are generated through cleavage at the anticodon. A given tRNA typically gives rise to several
[...] Read more.
Transfer RNA-derived fragments (tRFs) are noncoding RNAs that arise from either mature transfer RNAs (tRNAs) or their precursors. One important category of tRFs comprises the tRNA halves, which are generated through cleavage at the anticodon. A given tRNA typically gives rise to several co-expressed 5’-tRNA halves (5′-tRHs) that differ in the location of their 3′ ends. These 5′-tRHs, even though distinct, have traditionally been treated as indistinguishable from one another due to their near-identical sequences and lengths. We focused on co-expressed 5′-tRHs that arise from the same tRNA and systematically examined their exact sequences and abundances across 10 different human tissues. To this end, we manually curated and analyzed several hundred human RNA-seq datasets from NCBI’s Sequence Run Archive (SRA). We grouped datasets from the same tissue into their own collection and examined each group separately. We found that a given tRNA produces different groups of co-expressed 5′-tRHs in different tissues, different cell lines, and different diseases. Importantly, the co-expressed 5′-tRHs differ in their sequences, absolute abundances, and relative abundances, even among tRNAs with near-identical sequences from the same isodecoder or isoacceptor group. The findings suggest that co-expressed 5′-tRHs that are produced from the same tRNA or closely related tRNAs have distinct, context-dependent roles. Moreover, our analyses show that cell lines modeling the same tissue type and disease may not be interchangeable when it comes to experimenting with tRFs.
Full article
(This article belongs to the Special Issue Small RNAs – Big Roles: IsomiRs, tRNA Fragments, and rRNA Fragments in Human Health and Disease)
►▼
Show Figures
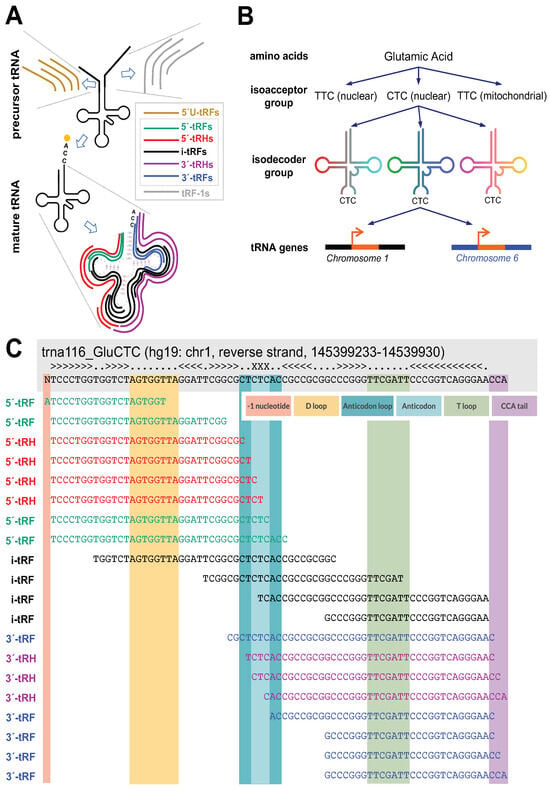
Figure 1
Open AccessReview
Dysregulation of Non-Coding RNAs: Roles of miRNAs and lncRNAs in the Pathogenesis of Multiple Myeloma
by
, , , , , , and
Non-Coding RNA 2023, 9(6), 68; https://doi.org/10.3390/ncrna9060068 - 03 Nov 2023
Abstract
The dysregulation of non-coding RNAs (ncRNAs), specifically microRNAs (miRNAs) and long non-coding RNAs (lncRNAs), leads to the development and advancement of multiple myeloma (MM). miRNAs, in particular, are paramount in post-transcriptional gene regulation, promoting mRNA degradation and translational inhibition. As a result, miRNAs
[...] Read more.
The dysregulation of non-coding RNAs (ncRNAs), specifically microRNAs (miRNAs) and long non-coding RNAs (lncRNAs), leads to the development and advancement of multiple myeloma (MM). miRNAs, in particular, are paramount in post-transcriptional gene regulation, promoting mRNA degradation and translational inhibition. As a result, miRNAs can serve as oncogenes or tumor suppressors depending on the target genes. In MM, miRNA disruption could result in abnormal gene expression responsible for cell growth, apoptosis, and other biological processes pertinent to cancer development. The dysregulated miRNAs inhibit the activity of tumor suppressor genes, contributing to disease progression. Nonetheless, several miRNAs are downregulated in MM and have been identified as gene regulators implicated in extracellular matrix remodeling and cell adhesion. miRNA depletion potentially facilitates the tumor advancement and resistance of therapeutic drugs. Additionally, lncRNAs are key regulators of numerous cellular processes, such as gene expression, chromatin remodeling, protein trafficking, and recently linked MM development. The lncRNAs are uniquely expressed and influence gene expression that supports MM growth, in addition to facilitating cellular proliferation and viability via multiple molecular pathways. miRNA and lncRNA alterations potentially result in anomalous gene expression and interfere with the regular functioning of MM. Thus, this review aims to highlight the dysregulation of these ncRNAs, which engender novel therapeutic modalities for the treatment of MM.
Full article
(This article belongs to the Special Issue Current Strategies to Investigate the Role of ncRNAs in Carcinogenesis)
►▼
Show Figures
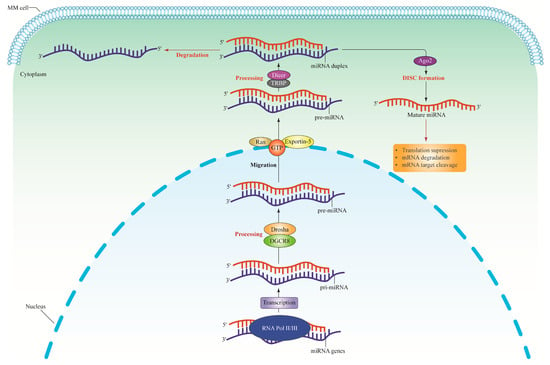
Figure 1
Open AccessArticle
Downregulation of Exosomal hsa-miR-551b-3p in Obesity and Its Link to Type 2 Diabetes Mellitus
by
, , , , , , , and
Non-Coding RNA 2023, 9(6), 67; https://doi.org/10.3390/ncrna9060067 - 02 Nov 2023
Abstract
Obesity is a significant risk factor for the development of type 2 diabetes mellitus (T2DM). Adipose tissue dysfunction can affect the pool of circulating exosomal miRNAs, driving concomitant disease in obesity. These exosomal miRNAs can reflect adipose tissue functionality, thus serving as prognostic
[...] Read more.
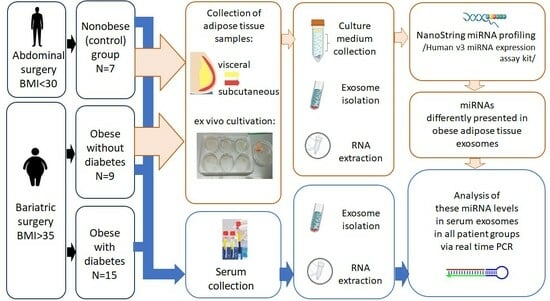
Obesity is a significant risk factor for the development of type 2 diabetes mellitus (T2DM). Adipose tissue dysfunction can affect the pool of circulating exosomal miRNAs, driving concomitant disease in obesity. These exosomal miRNAs can reflect adipose tissue functionality, thus serving as prognostic biomarkers for disease monitoring in case of T2DM. In the present study, we conducted NanoString microRNA profiling of extracellular vesicles (EVs) secreted by adipose tissue of obese patients (body mass index (BMI) > 35) without T2DM and nonobese individuals (BMI < 30) as a control group. Functional and pathway enrichment analysis showed that miRNAs associated with obesity in this study were implicated in insulin signaling and insulin resistance biological pathways. Further, these microRNAs were screened in serum EVs in the following groups: (1) obese patients with T2DM, (2) obese patients without T2DM, and (3) nonobese individuals as a control group. has-miR-551b-3p was shown to be downregulated in adipose tissue EVs, as well as in serum EVs, of patients with obesity without T2DM. At the same time, the serum exosomal hsa-miR-551b-3p content was significantly higher in obese patients with T2DM when compared with obese patients without T2DM and may be a potential biomarker of T2DM development in obesity.
Full article
(This article belongs to the Special Issue Non-coding RNA and Diabetes 2.0)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Organotypic 3D Cell-Architecture Impacts the Expression Pattern of miRNAs–mRNAs Network in Breast Cancer SKBR3 Cells
by
, , , , , , , , , and
Non-Coding RNA 2023, 9(6), 66; https://doi.org/10.3390/ncrna9060066 - 26 Oct 2023
Abstract
Background. Currently, most of the research on breast cancer has been carried out in conventional two-dimensional (2D) cell cultures due to its practical benefits, however, the three-dimensional (3D) cell culture is becoming the model of choice in cancer research because it allows cell–cell
[...] Read more.
Background. Currently, most of the research on breast cancer has been carried out in conventional two-dimensional (2D) cell cultures due to its practical benefits, however, the three-dimensional (3D) cell culture is becoming the model of choice in cancer research because it allows cell–cell and cell–extracellular matrix (ECM) interactions, mimicking the native microenvironment of tumors in vivo. Methods. In this work, we evaluated the effect of 3D cell organization on the expression pattern of miRNAs (by Small-RNAseq) and mRNAs (by microarrays) in the breast cancer SKBR3 cell line and analyzed the biological processes and signaling pathways regulated by the differentially expressed protein-coding genes (DE-mRNAs) and miRNAs (DE-microRNAs) found in the organoids. Results. We obtained well-defined cell-aggregated organoids with a grape cluster-like morphology with a size up to 9.2 × 105 μm3. The transcriptomic assays showed that cell growth in organoids significantly affected (all p < 0.01) the gene expression patterns of both miRNAs, and mRNAs, finding 20 upregulated and 19 downregulated DE-microRNAs, as well as 49 upregulated and 123 downregulated DE-mRNAs. In silico analysis showed that a subset of 11 upregulated DE-microRNAs target 70 downregulated DE-mRNAs. These genes are involved in 150 gene ontology (GO) biological processes such as regulation of cell morphogenesis, regulation of cell shape, regulation of canonical Wnt signaling pathway, morphogenesis of epithelium, regulation of cytoskeleton organization, as well as in the MAPK and AGE–RAGE signaling KEGG-pathways. Interestingly, hsa-mir-122-5p (Fold Change (FC) = 15.4), hsa-mir-369-3p (FC = 11.4), and hsa-mir-10b-5p (FC = 20.1) regulated up to 81% of the 70 downregulated DE-mRNAs. Conclusion. The organotypic 3D cell-organization architecture of breast cancer SKBR3 cells impacts the expression pattern of the miRNAs–mRNAs network mainly through overexpression of hsa-mir-122-5p, hsa-mir-369-3p, and hsa-mir-10b-5p. All these findings suggest that the interaction between cell–cell and cell–ECM as well as the change in the culture architecture impacts gene expression, and, therefore, support the pertinence of migrating breast cancer research from conventional cultures to 3D models.
Full article
(This article belongs to the Topic Non-coding RNAs in Cancer: Markers in Disease Progression and Therapy)
►▼
Show Figures
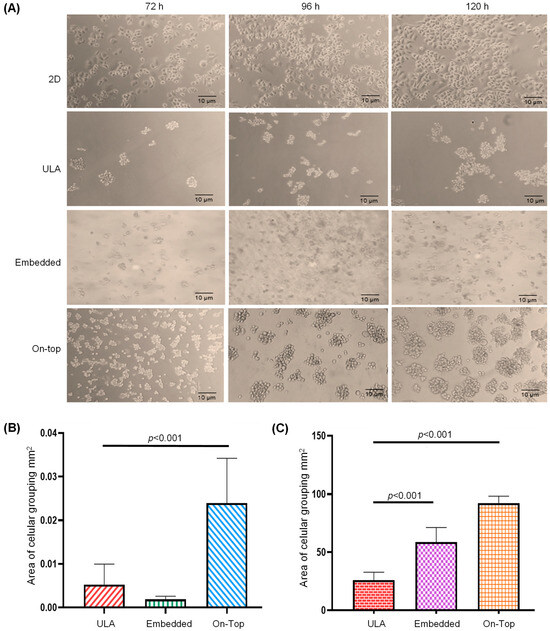
Figure 1
Open AccessArticle
Discovery of Novel miRNAs in Colorectal Cancer: Potential Biological Roles and Clinical Utility
by
, , , , , , , , , and
Non-Coding RNA 2023, 9(6), 65; https://doi.org/10.3390/ncrna9060065 - 26 Oct 2023
Abstract
Deregulated miRNAs are associated with colorectal cancer (CRC), with alterations depending on the tumor location. Novel tissue-specific miRNAs have been identified in different tumors and are associated with cancer. We used miRMaster to identify novel miRNAs in CRC from the TCGA and GEO
[...] Read more.
Deregulated miRNAs are associated with colorectal cancer (CRC), with alterations depending on the tumor location. Novel tissue-specific miRNAs have been identified in different tumors and are associated with cancer. We used miRMaster to identify novel miRNAs in CRC from the TCGA and GEO data (discovery and validation groups). We used TCGA data from five tissues to analyze miRNA tissue specificity. miRDB was used to predict miRNA targets, and the UCSC Xena Browser was used to evaluate target expression. After successive analyses, we identified 15 novel miRNAs with the same expression patterns in CRC in both the discovery and validation groups. Four molecules (nov-miR-13844-5p, nov-miR-7154-5p, nov-miR-5035-3p, and nov-miR-590-5p) were differentially expressed in proximal and distal CRC. The nov-miR-3345-5p and nov-miR-13172-3p, which are upregulated in tumors, are only expressed in colorectal tissues. These molecules have been linked to a worse prognosis in right-sided colon and rectal carcinomas. An analysis revealed an association between eight novel miRNAs and 81 targets, mostly cancer-related genes, with varying expression based on tumor location. These findings provide new miRNAs with potential biological relevance, molecular biomarkers, and therapeutic targets for CRC treatment.
Full article
(This article belongs to the Special Issue Molecular Mechanisms and Clinical Implications of Non-coding RNAs in Cancer)
►▼
Show Figures
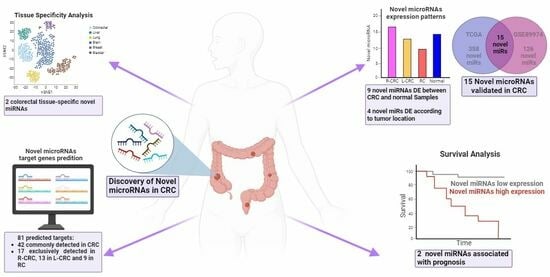
Graphical abstract
Open AccessArticle
A Machine Learning Model Based on microRNAs for the Diagnosis of Essential Hypertension
Non-Coding RNA 2023, 9(6), 64; https://doi.org/10.3390/ncrna9060064 - 25 Oct 2023
Abstract
Introduction: Hypertension is a major and modifiable risk factor for cardiovascular diseases. Essential, primary, or idiopathic hypertension accounts for 90–95% of all cases. Identifying novel biomarkers specific to essential hypertension may help in understanding pathophysiological pathways and developing personalized treatments. We tested whether
[...] Read more.
Introduction: Hypertension is a major and modifiable risk factor for cardiovascular diseases. Essential, primary, or idiopathic hypertension accounts for 90–95% of all cases. Identifying novel biomarkers specific to essential hypertension may help in understanding pathophysiological pathways and developing personalized treatments. We tested whether the integration of circulating microRNAs (miRNAs) and clinical risk factors via machine learning modeling may provide useful information and novel tools for essential hypertension diagnosis and management. Materials and methods: In total, 174 participants were enrolled in the present observational case–control study, among which, there were 89 patients with essential hypertension and 85 controls. A discovery phase was conducted using small RNA sequencing in whole blood samples obtained from age- and sex-matched hypertension patients (n = 30) and controls (n = 30). A validation phase using RT-qPCR involved the remaining 114 participants. For machine learning, 170 participants with complete data were used to generate and evaluate the classification model. Results: Small RNA sequencing identified seven miRNAs downregulated in hypertensive patients as compared with controls in the discovery group, of which six were confirmed with RT-qPCR. In the validation group, miR-210-3p/361-3p/362-5p/378a-5p/501-5p were also downregulated in hypertensive patients. A machine learning support vector machine (SVM) model including clinical risk factors (sex, BMI, alcohol use, current smoker, and hypertension family history), miR-361-3p, and miR-501-5p was able to classify hypertension patients in a test dataset with an AUC of 0.90, a balanced accuracy of 0.87, a sensitivity of 0.83, and a specificity of 0.91. While five miRNAs exhibited substantial downregulation in hypertension patients, only miR-361-3p and miR-501-5p, alongside clinical risk factors, were consistently chosen in at least eight out of ten sub-training sets within the SVM model. Conclusions: This study highlights the potential significance of miRNA-based biomarkers in deepening our understanding of hypertension’s pathophysiology and in personalizing treatment strategies. The strong performance of the SVM model highlights its potential as a valuable asset for diagnosing and managing essential hypertension. The model remains to be extensively validated in independent patient cohorts before evaluating its added value in a clinical setting.
Full article
(This article belongs to the Collection Regulatory RNAs in Cardiovascular Development and Disease)
►▼
Show Figures
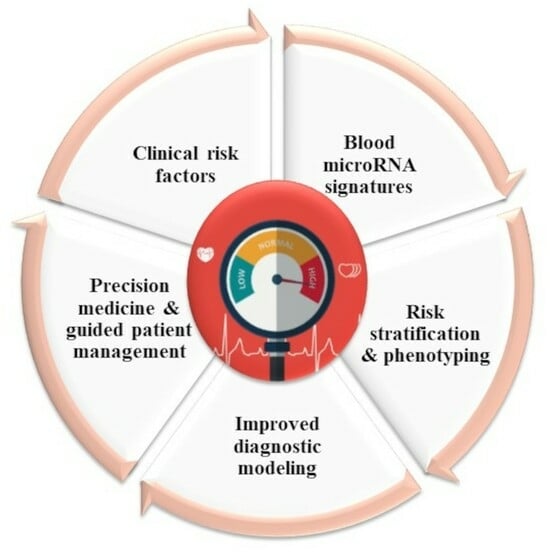
Graphical abstract
Open AccessReview
Crosstalk between Non-Coding RNAs and Wnt/β-Catenin Signaling in Head and Neck Cancer: Identification of Novel Biomarkers and Therapeutic Agents
by
, , , , , and
Non-Coding RNA 2023, 9(5), 63; https://doi.org/10.3390/ncrna9050063 - 17 Oct 2023
Abstract
Head and neck cancers (HNC) encompass a broad spectrum of neoplastic disorders characterized by significant morbidity and mortality. While contemporary therapeutic interventions offer promise, challenges persist due to tumor recurrence and metastasis. Central to HNC pathogenesis is the aberration in numerous signaling cascades.
[...] Read more.
Head and neck cancers (HNC) encompass a broad spectrum of neoplastic disorders characterized by significant morbidity and mortality. While contemporary therapeutic interventions offer promise, challenges persist due to tumor recurrence and metastasis. Central to HNC pathogenesis is the aberration in numerous signaling cascades. Prominently, the Wnt signaling pathway has been critically implicated in the etiology of HNC, as supported by a plethora of research. Equally important, variations in the expression of non-coding RNAs (ncRNAs) have been identified to modulate key cancer phenotypes such as cellular proliferation, epithelial-mesenchymal transition, metastatic potential, recurrence, and treatment resistance. This review aims to provide an exhaustive insight into the multifaceted influence of ncRNAs on HNC, with specific emphasis on their interactions with the Wnt/β-catenin (WBC) signaling axis. We further delineate the effect of ncRNAs in either exacerbating or attenuating HNC progression via interference with WBC signaling. An overview of the mechanisms underlying the interplay between ncRNAs and WBC signaling is also presented. In addition, we described the potential of various ncRNAs in enhancing the efficacy of chemotherapeutic and radiotherapeutic modalities. In summary, this assessment posits the potential of ncRNAs as therapeutic agents targeting the WBC signaling pathway in HNC management.
Full article
(This article belongs to the Special Issue Molecular Mechanisms and Clinical Implications of Non-coding RNAs in Cancer)
►▼
Show Figures
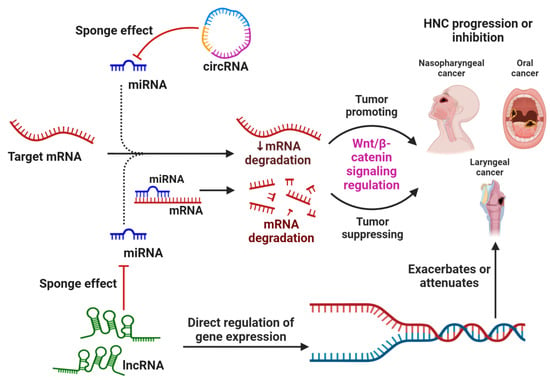
Figure 1
Open AccessArticle
The Potential Role of Circulating Long Miscellaneous RNAs in the Diagnosis and Prognosis of Hepatitis C Related Hepatocellular Carcinoma
by
, , , , , , , , , , , and
Non-Coding RNA 2023, 9(5), 62; https://doi.org/10.3390/ncrna9050062 - 11 Oct 2023
Abstract
►▼
Show Figures
Ribonucleic acids (RNAs) are important regulators of gene expression and crucial for the progression of hepatocellular carcinoma (HCC). This study was designed to determine the diagnostic and prognostic utility of the circulating long miscellaneous RNAs; LINC01419, AK021443, and AF070632 in HCV-related HCC patients.
[...] Read more.
Ribonucleic acids (RNAs) are important regulators of gene expression and crucial for the progression of hepatocellular carcinoma (HCC). This study was designed to determine the diagnostic and prognostic utility of the circulating long miscellaneous RNAs; LINC01419, AK021443, and AF070632 in HCV-related HCC patients. Real-time PCR was used to measure their relative expression levels in the plasma of 194 HCV patients, 120 HCV-related HCC patients and 120 healthy controls. LINC01419 and AK021443 expression levels had significantly increasing linear trend estimates while AF070632 was dramatically downregulated in HCC compared to HCV. Interestingly, LINC01419 and AK021443 served as more significant diagnostic biomarkers for HCC than AF070632 and AFP. Multivariate analysis with cox regression revealed that the high expression of AK021443 [HR = 10.06, CI95%: 3.36–30.07], the high expression of LINC01419 [HR 4.13, CI95%: 1.32–12.86], and the low expression of AF070632 [HR = 2.70, CI95%: 1.07–6.81] were significant potential prognostic factors for HCC. Besides, the Kaplan–Meier analysis showed that HCC patients with high LIN01419 and AK021443 and low AF070632 expression levels had shorter OS. The circulating LINC01419 and AK021443 can be used as noninvasive potential biomarkers for diagnosis and prognosis of HCV-related HCC patients than AF070632 providing new targets for limiting the progression of the disease.
Full article

Figure 1

Journal Menu
► ▼ Journal Menu-
- ncRNA Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Topical Collections
- Article Processing Charge
- Indexing & Archiving
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomolecules, Cells, Genes, IJMS, ncRNA
MicroRNA: Mechanisms of Action, Physio-Pathological Implications, and Disease Biomarkers, 2nd Volume
Topic Editors: Y-h. Taguchi, Hsiuying WangDeadline: 20 April 2024

Conferences
Special Issues
Special Issue in
ncRNA
Non-coding RNAs and Phase Separation Phenomena
Guest Editors: Francisco J. Enguita, Ana Lúcia LeitãoDeadline: 28 February 2024
Special Issue in
ncRNA
Roles of Non-coding RNAs in Drug Metabolism and Disposition
Guest Editors: Xiaobo Zhong, Lirong ZhangDeadline: 15 March 2024
Special Issue in
ncRNA
ncRNA and Cancer Immunotherapy
Guest Editor: Agnieszka M. BroniszDeadline: 31 March 2024
Special Issue in
ncRNA
circRNAs in Human Healthy and Diseases
Guest Editor: Xiao-Ou ZhangDeadline: 20 April 2024
Topical Collections
Topical Collection in
ncRNA
Secreted Non-coding RNAs as Signaling Molecules Driving Cell-to-Cell Communication in Cancer
Collection Editor: Giulia Fontemaggi
Topical Collection in
ncRNA
Non-coding RNAs, COVID-19, and Long-COVID
Collection Editors: Gaetano Santulli, Jessica Gambardella
Topical Collection in
ncRNA
The Non-Coding RNA Journal Club: Highlights on Recent Papers
Collection Editor: George A. Calin









_Cropped_2.jpg)














