-
 Respiratory Syncytial Virus Vaccines: A Review of the Candidates and the Approved Vaccines
Respiratory Syncytial Virus Vaccines: A Review of the Candidates and the Approved Vaccines -
 The Cultured Microbiome of Pollinated Maize Silks Shifts after Infection with Fusarium graminearum and Varies by Distance from the Site of Pathogen Inoculation
The Cultured Microbiome of Pollinated Maize Silks Shifts after Infection with Fusarium graminearum and Varies by Distance from the Site of Pathogen Inoculation -
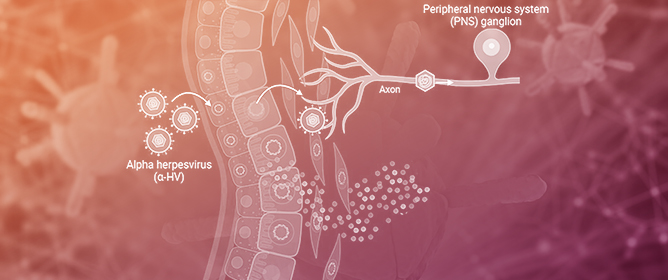 Interferon-λ Activates a Differential Response in Peripheral Neurons That Is Effective against Alpha Herpesvirus Infections
Interferon-λ Activates a Differential Response in Peripheral Neurons That Is Effective against Alpha Herpesvirus Infections
Journal Description
Pathogens
Pathogens
is an international, peer-reviewed, open access journal on pathogens and pathogen-host interactions published monthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, Embase, PubAg, CaPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (General Immunology and Microbiology)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.4 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Pathogens include: Parasitologia, Bacteria and Zoonotic Diseases.
Impact Factor:
3.7 (2022);
5-Year Impact Factor:
3.7 (2022)
Latest Articles
Mycoplasma gallisepticum and Mycoplasma synoviae in Turkeys in Poland
Pathogens 2024, 13(1), 78; https://doi.org/10.3390/pathogens13010078 - 15 Jan 2024
Abstract
The pathogenic mycoplasmas are among the bacteria causing significant losses in the poultry industry worldwide. Mycoplasma gallisepticum (MG) and M. synoviae (MS) are economically important pathogens causing chronic respiratory disease, decreased growth, egg production and hatchability rates, and significant downgrading of carcasses. Effective
[...] Read more.
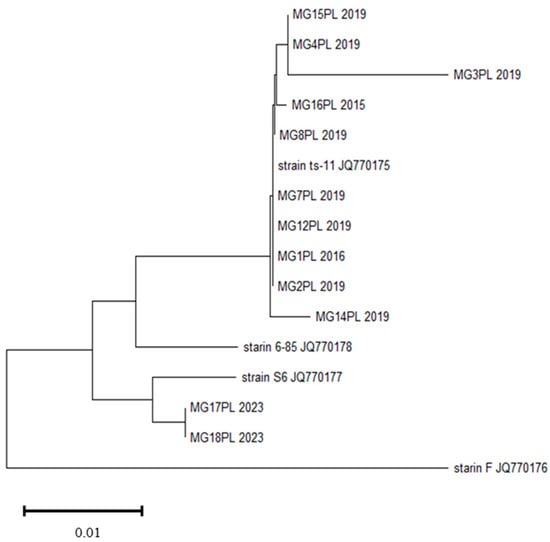
The pathogenic mycoplasmas are among the bacteria causing significant losses in the poultry industry worldwide. Mycoplasma gallisepticum (MG) and M. synoviae (MS) are economically important pathogens causing chronic respiratory disease, decreased growth, egg production and hatchability rates, and significant downgrading of carcasses. Effective diagnosis of infection with these species in poultry is highly requisite considering their two routes of spreading—horizontal and vertical. Their prevalence and molecular epidemiology were investigated in 184 turkey flocks in Poland. Tracheal samples were selected from 144 broiler flocks and 40 turkey breeder flocks collected in 2015–2023. The prevalence of MG was determined by real-time PCR targeting the 16S rRNA gene and PCR targeting the mgc2 gene, and MS was determined by a 16–23S rRNA real-time PCR and a vlhA gene PCR. Further identification and molecular characterization were carried out using PCR and sequencing. M. gallisepticum and M. synoviae were found in 8.33% and 9.72% of turkey broiler flocks respectively. The phylogenetic analysis of MG isolates in most cases showed high similarity to the ts-11-like strains. MS isolates showed high similarity to strains isolated from flocks of laying hens causing EAA. Additional tests detected Ornithobacterium rhinotracheale, Gallibacterium anatis, Enterococcus faecalis and Enterococcus faecium, Staphylococcus aureus and Riemerella anatipestifer. These secondary pathogens could have significantly heightened the pathogenicity of the mycoplasma infections studied.
Full article
(This article belongs to the Section Bacterial Pathogens)
►
Show Figures
Open AccessArticle
First Report of Benzimidazole Resistance in Field Population of Haemonchus contortus from Sheep, Goats and Cattle in Bosnia and Herzegovina
by
, , , , , , , and
Pathogens 2024, 13(1), 77; https://doi.org/10.3390/pathogens13010077 - 15 Jan 2024
Abstract
Haemonchus contortus is a globally significant parasitic nematode in ruminants, with widespread resistance to benzimidazole due to its excessive and prolonged use. Given the extensive use of benzimidazole anthelmintics in Bosnia and Herzegovina, we hypothesized that resistance is prevalent. The aim of this
[...] Read more.
Haemonchus contortus is a globally significant parasitic nematode in ruminants, with widespread resistance to benzimidazole due to its excessive and prolonged use. Given the extensive use of benzimidazole anthelmintics in Bosnia and Herzegovina, we hypothesized that resistance is prevalent. The aim of this study was to identify the presence of anthelmintic resistance to benzimidazole in H. contortus from naturally infected sheep, goats and cattle in Bosnia and Herzegovina through the detection of the Phe/Tyr polymorphism in the amino acid at position 200 of the β-tubulin protein. From 19 locations in Bosnia and Herzegovina, a total of 83 adult H. contortus were collected from the abomasum of ruminants. Among these, 45 H. contortus specimens were isolated from sheep, 19 from goats and 19 from cattle. Results showed that 77.8% of H. contortus in sheep exhibited homozygous resistant genotypes at position 200 of the β-tubulin gene, with 15.5% being heterozygous. In goats, all tested H. contortus (100%) were homozygous resistant, and no heterozygous resistant or homozygous sensitive genotypes were found. Cattle had 94.7% homozygous resistant H. contortus, with no heterozygous resistant genotypes detected. In H. contortus from sheep and cattle, 6.7% and 5.3%, respectively, displayed homozygous sensitive genotypes. This study, for the first time, highlights the presence of a resistant population of H. contortus in sheep, goats and cattle in Bosnia and Herzegovina, using the rt-qPCR method. The resistance likely spread from sheep or goats to cattle, facilitated by shared pastures and the practice of transhumance, indicating a widespread and growing issue of anthelmintic resistance.
Full article
(This article belongs to the Special Issue Parasitic Helminths: Drug Resistance, Control and Immune Evasion)
►▼
Show Figures
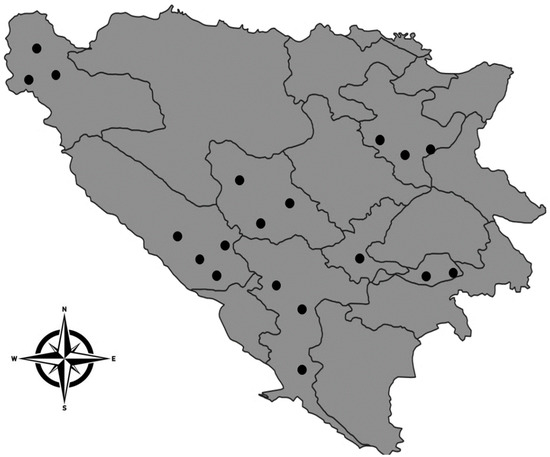
Figure 1
Open AccessReview
Biofilm Producing Methicillin-Resistant Staphylococcus aureus (MRSA) Infections in Humans: Clinical Implications and Management
Pathogens 2024, 13(1), 76; https://doi.org/10.3390/pathogens13010076 - 15 Jan 2024
Abstract
Since its initial description in the 1960s, methicillin-resistant Staphylococcus aureus (MRSA) has developed multiple mechanisms for antimicrobial resistance and evading the immune system, including biofilm production. MRSA is now a widespread pathogen, causing a spectrum of infections ranging from superficial skin issues to
[...] Read more.
Since its initial description in the 1960s, methicillin-resistant Staphylococcus aureus (MRSA) has developed multiple mechanisms for antimicrobial resistance and evading the immune system, including biofilm production. MRSA is now a widespread pathogen, causing a spectrum of infections ranging from superficial skin issues to severe conditions like osteoarticular infections and endocarditis, leading to high morbidity and mortality. Biofilm production is a key aspect of MRSA’s ability to invade, spread, and resist antimicrobial treatments. Environmental factors, such as suboptimal antibiotics, pH, temperature, and tissue oxygen levels, enhance biofilm formation. Biofilms are intricate bacterial structures with dense organisms embedded in polysaccharides, promoting their resilience. The process involves stages of attachment, expansion, maturation, and eventually disassembly or dispersion. MRSA’s biofilm formation has a complex molecular foundation, involving genes like icaADBC, fnbA, fnbB, clfA, clfB, atl, agr, sarA, sarZ, sigB, sarX, psm, icaR, and srtA. Recognizing pivotal genes for biofilm formation has led to potential therapeutic strategies targeting elemental and enzymatic properties to combat MRSA biofilms. This review provides a practical approach for healthcare practitioners, addressing biofilm pathogenesis, disease spectrum, and management guidelines, including advances in treatment. Effective management involves appropriate antimicrobial therapy, surgical interventions, foreign body removal, and robust infection control practices to curtail spread within healthcare environments.
Full article
(This article belongs to the Special Issue Advances in Treatment of Biofilm Infections)
Open AccessReview
SARS-CoV-2 ORF3a Protein as a Therapeutic Target against COVID-19 and Long-Term Post-Infection Effects
by
, , , , , , , , and
Pathogens 2024, 13(1), 75; https://doi.org/10.3390/pathogens13010075 - 14 Jan 2024
Abstract
The COVID-19 pandemic caused by SARS-CoV-2 has posed unparalleled challenges due to its rapid transmission, ability to mutate, high mortality and morbidity, and enduring health complications. Vaccines have exhibited effectiveness, but their efficacy diminishes over time while new variants continue to emerge. Antiviral
[...] Read more.
The COVID-19 pandemic caused by SARS-CoV-2 has posed unparalleled challenges due to its rapid transmission, ability to mutate, high mortality and morbidity, and enduring health complications. Vaccines have exhibited effectiveness, but their efficacy diminishes over time while new variants continue to emerge. Antiviral medications offer a viable alternative, but their success has been inconsistent. Therefore, there remains an ongoing need to identify innovative antiviral drugs for treating COVID-19 and its post-infection complications. The ORF3a (open reading frame 3a) protein found in SARS-CoV-2, represents a promising target for antiviral treatment due to its multifaceted role in viral pathogenesis, cytokine storms, disease severity, and mortality. ORF3a contributes significantly to viral pathogenesis by facilitating viral assembly and release, essential processes in the viral life cycle, while also suppressing the body’s antiviral responses, thus aiding viral replication. ORF3a also has been implicated in triggering excessive inflammation, characterized by NF-κB-mediated cytokine production, ultimately leading to apoptotic cell death and tissue damage in the lungs, kidneys, and the central nervous system. Additionally, ORF3a triggers the activation of the NLRP3 inflammasome, inciting a cytokine storm, which is a major contributor to the severity of the disease and subsequent mortality. As with the spike protein, ORF3a also undergoes mutations, and certain mutant variants correlate with heightened disease severity in COVID-19. These mutations may influence viral replication and host cellular inflammatory responses. While establishing a direct link between ORF3a and mortality is difficult, its involvement in promoting inflammation and exacerbating disease severity likely contributes to higher mortality rates in severe COVID-19 cases. This review offers a comprehensive and detailed exploration of ORF3a’s potential as an innovative antiviral drug target. Additionally, we outline potential strategies for discovering and developing ORF3a inhibitor drugs to counteract its harmful effects, alleviate tissue damage, and reduce the severity of COVID-19 and its lingering complications.
Full article
(This article belongs to the Section Vaccines and Therapeutic Developments)
►▼
Show Figures
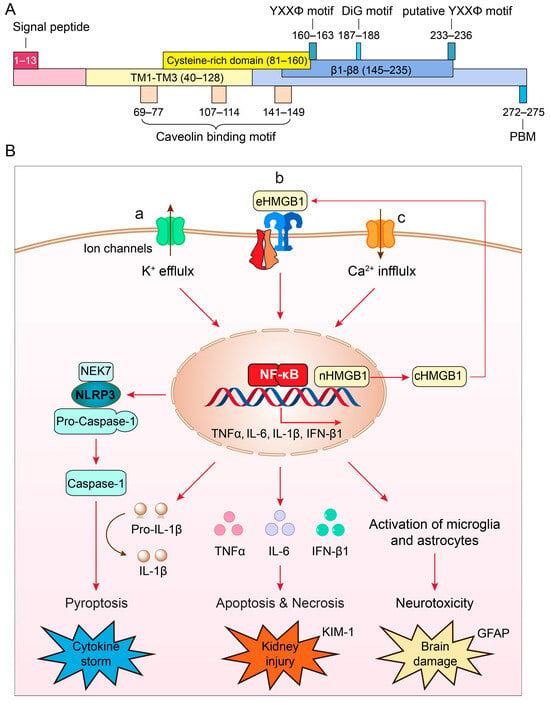
Figure 1
Open AccessArticle
Complement Suppresses the Initial Type 1 Interferon Response to Ocular Herpes Simplex Virus Type 1 Infection in Mice
Pathogens 2024, 13(1), 74; https://doi.org/10.3390/pathogens13010074 - 13 Jan 2024
Abstract
The complement system (CS) contributes to the initial containment of viral and bacterial pathogens and clearance of dying cells in circulation. We previously reported mice deficient in complement component 3 (C3KO mice) were more sensitive than wild-type (WT) mice to ocular HSV-1 infection,
[...] Read more.
The complement system (CS) contributes to the initial containment of viral and bacterial pathogens and clearance of dying cells in circulation. We previously reported mice deficient in complement component 3 (C3KO mice) were more sensitive than wild-type (WT) mice to ocular HSV-1 infection, as measured by a reduction in cumulative survival and elevated viral titers in the nervous system but not the cornea between days three and seven post infection (pi). The present study was undertaken to determine if complement deficiency impacted virus replication and associated changes in inflammation at earlier time points in the cornea. C3KO mice were found to possess significantly (p < 0.05) less infectious virus in the cornea at 24 h pi that corresponded with a decrease in HSV-1 lytic gene expression at 12 and 24 h pi compared to WT animals. Flow cytometry acquisition found no differences in the myeloid cell populations residing in the cornea including total macrophage and neutrophil populations at 24 h pi with minimal infiltrating cell populations detected at the 12 h pi time point. Analysis of cytokine and chemokine content in the cornea measured at 12 and 24 h pi revealed that only CCL3 (MIP-1α) was found to be different between WT and C3KO mice with >2-fold increased levels (p < 0.05, ANOVA and Tukey’s post hoc t-test) in the cornea of WT mice at 12 h pi. C3KO mouse resistance to HSV-1 infection at the early time points correlated with a significant increase in type I interferon (IFN) gene expression including IFN-α1 and IFN-β and downstream effector genes including tetherin and RNase L (p < 0.05, Mann–Whitney rank order test). These results suggest early activation of the CS interferes with the induction of the type I IFN response and leads to a transient increase in virus replication following corneal HSV-1 infection.
Full article
(This article belongs to the Special Issue Editorial Board Members’ Collection Series: Molecular Epidemiology, Diagnostics and Management of Viral Pathogens)
►▼
Show Figures
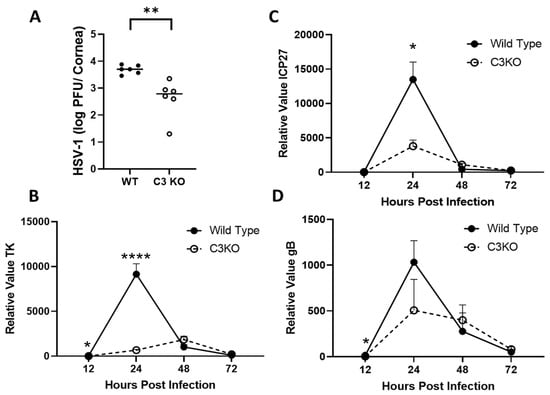
Figure 1
Open AccessEditorial
Biting Back: Advances in Fighting Ticks and Understanding Tick-Borne Pathogens
Pathogens 2024, 13(1), 73; https://doi.org/10.3390/pathogens13010073 - 13 Jan 2024
Abstract
Ticks are blood-feeding arthropods and obligate ectoparasites of virtually all animal species (except fish) and humans [...]
Full article
Open AccessArticle
Myxoma Virus Combination Therapy Enhances Lenalidomide and Bortezomib Treatments for Multiple Myeloma
by
, , , , , , , and
Pathogens 2024, 13(1), 72; https://doi.org/10.3390/pathogens13010072 - 12 Jan 2024
Abstract
This study aimed to explore the effectiveness and safety of Myxoma virus (MYXV) in MM cell lines and primary myeloma cells obtained from patients with multiple myeloma. Myeloma cells were isolated from MM patients and cultured. MYXV, lenalidomide, and bortezomib were used in
[...] Read more.
This study aimed to explore the effectiveness and safety of Myxoma virus (MYXV) in MM cell lines and primary myeloma cells obtained from patients with multiple myeloma. Myeloma cells were isolated from MM patients and cultured. MYXV, lenalidomide, and bortezomib were used in MM cells. The cytotoxicity assay was investigated using WST-1. Apoptosis was assessed through flow cytometry with Annexin V/PI staining and caspase-9 concentrations using ELISA. To explore MYXV entry into MM cells, monoclonal antibodies were used. Moreover, to explore the mechanisms of MYXV entry into MM cells, we examined the level of GFP-labeled MYXV within the cells after blocking with monoclonal antibodies targeting BCMA, CD20, CD28, CD33, CD38, CD56, CD86, CD117, CD138, CD200, and CD307 in MM cells. The study demonstrated the effects of treating Myxoma virus with lenalidomide and bortezomib. The treatment resulted in reduced cell viability and increased caspase-9 expression. Only low-dose CD86 blockade showed a significant difference in MYXV entry into MM cells. The virus caused an increase in the rate of apoptosis in the cells, regardless of whether it was administered alone or in combination with drugs. The groups with the presence of the virus showed higher rates of early apoptosis. The Virus, Virus + Bortezomib, and Virus + Lenalidomide groups had significantly higher rates of early apoptosis (p < 0.001). However, the measurements of late apoptosis and necrosis showed variability. The addition of MYXV resulted in a statistically significant increase in early apoptosis in both newly diagnosed and refractory MM patients. Our results highlight that patient-based therapy should also be considered for the effective management of MM.
Full article
(This article belongs to the Section Vaccines and Therapeutic Developments)
Open AccessArticle
A Genome-Wide Association Study for Resistance to Tropical Theileriosis in Two Bovine Portuguese Autochthonous Breeds
by
, , , , , , and
Pathogens 2024, 13(1), 71; https://doi.org/10.3390/pathogens13010071 - 12 Jan 2024
Abstract
The control of Tropical Theileriosis, a tick-borne disease with a strong impact on cattle breeding, can be facilitated using marker-assisted selection in breeding programs. Genome-wide association studies (GWAS) using high-density arrays are extremely important for the ongoing process of identifying genomic variants associated
[...] Read more.
The control of Tropical Theileriosis, a tick-borne disease with a strong impact on cattle breeding, can be facilitated using marker-assisted selection in breeding programs. Genome-wide association studies (GWAS) using high-density arrays are extremely important for the ongoing process of identifying genomic variants associated with resistance to Theileria annulata infection. In this work, single-nucleotide polymorphisms (SNPs) were analyzed in the Portuguese autochthonous cattle breeds Alentejana and Mertolenga. In total, 24 SNPs suggestive of significance (p ≤ 10−4) were identified for Alentejana cattle and 20 SNPs were identified for Mertolenga cattle. The genomic regions around these SNPs were further investigated for annotated genes and quantitative trait loci (QTLs) previously described by other authors. Regarding the Alentejana breed, the MAP3K1, CMTM7, SSFA2, and ATG13 genes are located near suggestive SNPs and appear as candidate genes for resistance to Tropical Theileriosis, considering its action in the immune response and resistance to other diseases. On the other hand, in the Mertolenga breed, the UOX gene is also a candidate gene due to its apparent link to the pathogenesis of the disease. These results may represent a first step toward the possibility of including genetic markers for resistance to Tropical Theileriosis in current breed selection programs.
Full article
(This article belongs to the Section Epidemiology of Infectious Diseases)
►▼
Show Figures
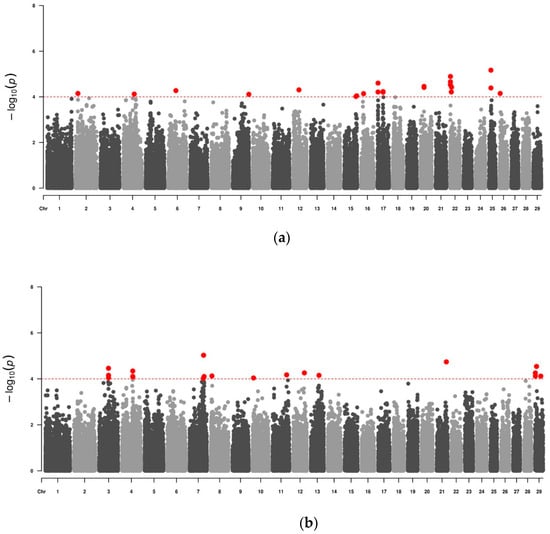
Figure 1
Open AccessArticle
Chromosome-Level Assemblies for the Pine Pitch Canker Pathogen Fusarium circinatum
by
, , , , , and
Pathogens 2024, 13(1), 70; https://doi.org/10.3390/pathogens13010070 - 12 Jan 2024
Abstract
The pine pitch canker pathogen, Fusarium circinatum, is globally regarded as one of the most important threats to commercial pine-based forestry. Although genome sequences of this fungus are available, these remain highly fragmented or structurally ill-defined. Our overall goal was to provide
[...] Read more.
The pine pitch canker pathogen, Fusarium circinatum, is globally regarded as one of the most important threats to commercial pine-based forestry. Although genome sequences of this fungus are available, these remain highly fragmented or structurally ill-defined. Our overall goal was to provide high-quality assemblies for two notable strains of F. circinatum, and to characterize these in terms of coding content, repetitiveness and the position of telomeres and centromeres. For this purpose, we used Oxford Nanopore Technologies MinION long-read sequences, as well as Illumina short sequence reads. By leveraging the genomic synteny inherent to F. circinatum and its close relatives, these sequence reads were assembled to chromosome level, where contiguous sequences mostly spanned from telomere to telomere. Comparative analyses unveiled remarkable variability in the twelfth and smallest chromosome, which is known to be dispensable. It presented a striking length polymorphism, with one strain lacking substantial portions from the chromosome’s distal and proximal regions. These regions, characterized by a lower gene density, G+C content and an increased prevalence of repetitive elements, contrast starkly with the syntenic segments of the chromosome, as well as with the core chromosomes. We propose that these unusual regions might have arisen or expanded due to the presence of transposable elements. A comparison of the overall chromosome structure revealed that centromeric elements often underpin intrachromosomal differences between F. circinatum strains, especially at chromosomal breakpoints. This suggests a potential role for centromeres in shaping the chromosomal architecture of F. circinatum and its relatives. The publicly available genome data generated here, together with the detailed metadata provided, represent essential resources for future studies of this important plant pathogen.
Full article
(This article belongs to the Special Issue Plant Pathogenic Fungi)
►▼
Show Figures
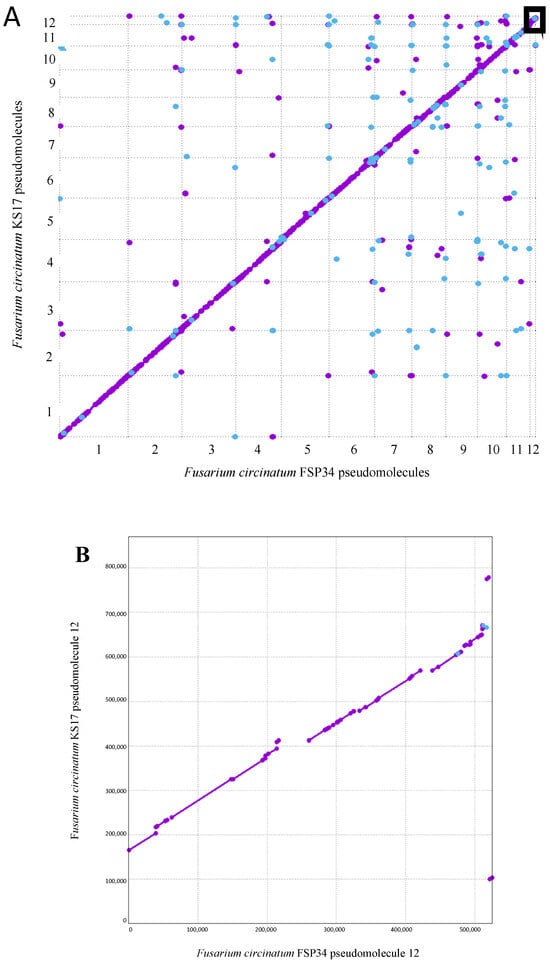
Figure 1
Open AccessReview
From Stress Tolerance to Virulence: Recognizing the Roles of Csps in Pathogenicity and Food Contamination
by
and
Pathogens 2024, 13(1), 69; https://doi.org/10.3390/pathogens13010069 - 11 Jan 2024
Abstract
Be it for lab studies or real-life situations, bacteria are constantly exposed to a myriad of physical or chemical stresses that selectively allow the tolerant to survive and thrive. In response to environmental fluctuations, the expression of cold shock domain family proteins (Csps)
[...] Read more.
Be it for lab studies or real-life situations, bacteria are constantly exposed to a myriad of physical or chemical stresses that selectively allow the tolerant to survive and thrive. In response to environmental fluctuations, the expression of cold shock domain family proteins (Csps) significantly increases to counteract and help cells deal with the harmful effects of stresses. Csps are, therefore, considered stress adaptation proteins. The primary functions of Csps include chaperoning nucleic acids and regulating global gene expression. In this review, we focus on the phenotypic effects of Csps in pathogenic bacteria and explore their involvement in bacterial pathogenesis. Current studies of csp deletions among pathogenic strains indicate their involvement in motility, host invasion and stress tolerance, proliferation, cell adhesion, and biofilm formation. Through their RNA chaperone activity, Csps regulate virulence-associated genes and thereby contribute to bacterial pathogenicity. Additionally, we outline their involvement in food contamination and discuss how foodborne pathogens utilize the stress tolerance roles of Csps against preservation and sanitation strategies. Furthermore, we highlight how Csps positively and negatively impact pathogens and the host. Overall, Csps are involved in regulatory networks that influence the expression of genes central to stress tolerance and virulence.
Full article
(This article belongs to the Section Bacterial Pathogens)
►▼
Show Figures
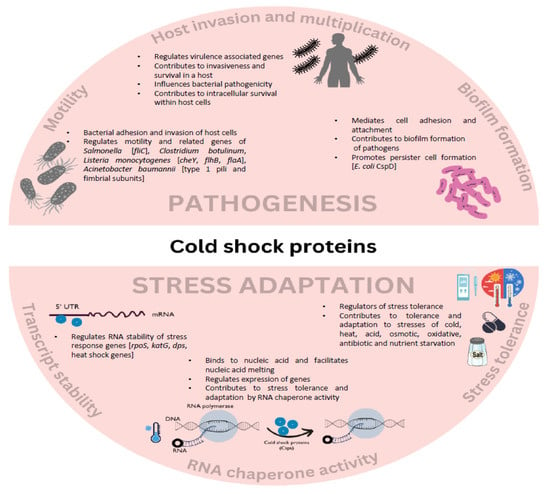
Figure 1
Open AccessReview
Metabolic Dysfunction-Associated Fatty Liver Disease and Chronic Viral Hepatitis: The Interlink
Pathogens 2024, 13(1), 68; https://doi.org/10.3390/pathogens13010068 - 10 Jan 2024
Abstract
Metabolic dysfunction-associated fatty liver disease (MAFLD) has now affected nearly one-third of the global population and has become the number one cause of chronic liver disease in the world because of the obesity pandemic. Chronic hepatitis resulting from hepatitis B virus (HBV) and
[...] Read more.
Metabolic dysfunction-associated fatty liver disease (MAFLD) has now affected nearly one-third of the global population and has become the number one cause of chronic liver disease in the world because of the obesity pandemic. Chronic hepatitis resulting from hepatitis B virus (HBV) and hepatitis C virus (HCV) remain significant challenges to liver health even in the 21st century. The co-existence of MAFLD and chronic viral hepatitis can markedly alter the disease course of individual diseases and can complicate the management of each of these disorders. A thorough understanding of the pathobiological interactions between MAFLD and these two chronic viral infections is crucial for appropriately managing these patients. In this comprehensive clinical review, we discuss the various mechanisms of chronic viral hepatitis-mediated metabolic dysfunction and the impact of MAFLD on the progression of liver disease.
Full article
(This article belongs to the Special Issue Viral Hepatitis in Europe: The Unresolved Issues)
►▼
Show Figures
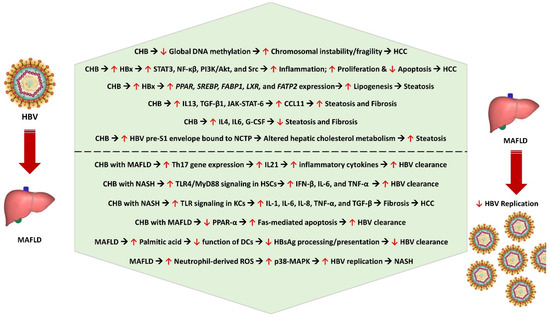
Figure 1
Open AccessArticle
Paracrine Signaling Mediated by the Cytosolic Tryparedoxin Peroxidase of Trypanosoma cruzi
by
, , , , , , and
Pathogens 2024, 13(1), 67; https://doi.org/10.3390/pathogens13010067 - 10 Jan 2024
Abstract
Peroxiredoxins are abundant and ubiquitous proteins that participate in different cellular functions, such as oxidant detoxification, protein folding, and intracellular signaling. Under different cellular conditions, peroxiredoxins can be secreted by different parasites, promoting the induction of immune responses in hosts. In this work,
[...] Read more.
Peroxiredoxins are abundant and ubiquitous proteins that participate in different cellular functions, such as oxidant detoxification, protein folding, and intracellular signaling. Under different cellular conditions, peroxiredoxins can be secreted by different parasites, promoting the induction of immune responses in hosts. In this work, we demonstrated that the cytosolic tryparedoxin peroxidase of Trypanosoma cruzi (cTXNPx) is secreted by epimastigotes and trypomastigotes associated with extracellular vesicles and also as a vesicle-free protein. By confocal microscopy, we show that cTXNPx can enter host cells by an active mechanism both through vesicles and as a recombinant protein. Transcriptomic analysis revealed that cTXNPx induces endoplasmic reticulum stress and interleukin-8 expression in epithelial cells. This analysis also suggested alterations in cholesterol metabolism in cTXNPx-treated cells, which was confirmed by immunofluorescence showing the accumulation of LDL and the induction of LDL receptors in both epithelial cells and macrophages. BrdU incorporation assays and qPCR showed that cTXNPx has a mitogenic, proliferative, and proinflammatory effect on these cells in a dose–dependent manner. Importantly, we also demonstrated that cTXNPx acts as a paracrine virulence factor, increasing the susceptibility to infection in cTXNPx-pretreated epithelial cells by approximately 40%. Although the results presented in this work are from in vitro studies and likely underestimate the complexity of parasite–host interactions, our work suggests a relevant role for this protein in establishing infection.
Full article
(This article belongs to the Section Parasitic Pathogens)
►▼
Show Figures
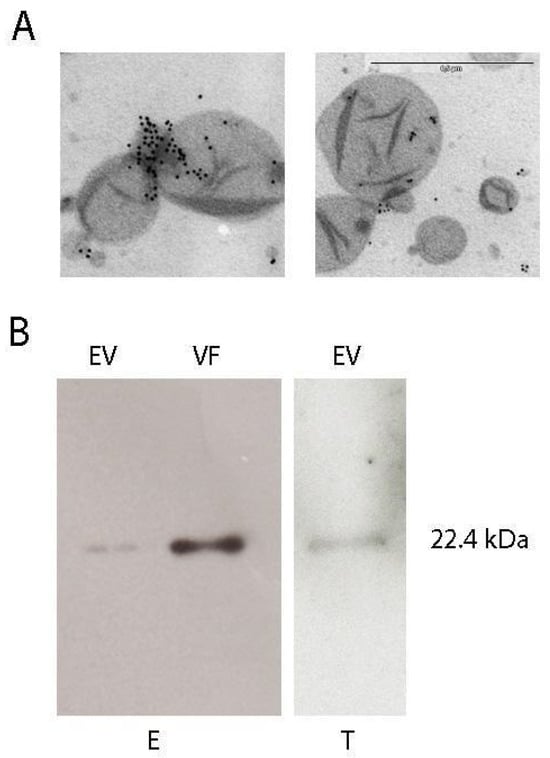
Figure 1
Open AccessCommunication
A Study of Cross-Protection between Eimeria maxima Immunovariants
Pathogens 2024, 13(1), 66; https://doi.org/10.3390/pathogens13010066 - 09 Jan 2024
Abstract
For reasons unknown, Eimeria maxima is unique among Eimeria species infecting chickens in the immunovariability it displays among isolates from different geographical areas. Eimeria maxima oocysts (named EmaxAPU3) were isolated late in grow-out (6 weeks) from litter in a commercial broiler operation that
[...] Read more.
For reasons unknown, Eimeria maxima is unique among Eimeria species infecting chickens in the immunovariability it displays among isolates from different geographical areas. Eimeria maxima oocysts (named EmaxAPU3) were isolated late in grow-out (6 weeks) from litter in a commercial broiler operation that was using Eimeria vaccination as the coccidiosis control program. Cross-protection studies (n = 4) were conducted in immunologically naïve chickens between EmaxAPU3 and two E. maxima lab strains (EmaxAPU1, EmaxAPU2) by immunizing with one E. maxima strain and challenging with either the homologous or heterologous E. maxima. As measured by oocyst output, immunization with EmaxAPU1 protected against homologous challenge (EmaxAPU1) and against heterologous challenge with EmaxAPU3, but not against EmaxAPU2. Similarly, immunization with EmaxAPU3 protected against homologous challenge (EmaxAPU3) and against heterologous challenge with EmaxAPU1, but not against EmaxAPU2. Immunization of chickens with EmaxAPU2 elicited a protective response against homologous challenge (EmaxAPU2), but not against EmaxAPU1 nor EmaxAPU3. The most plausible explanation for the appearance of this immunovariant late in grow-out is that E. maxima APU3 escaped immunity directed to E. maxima antigenic types in the commercial vaccine.
Full article
(This article belongs to the Special Issue Research on Coccidian Parasites in Livestock)
►▼
Show Figures
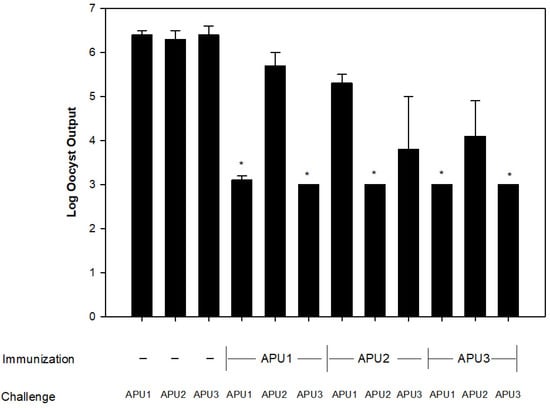
Figure 1
Open AccessEditorial
When Cell-Mediated Immunity after Vaccination Is Important
Pathogens 2024, 13(1), 65; https://doi.org/10.3390/pathogens13010065 - 09 Jan 2024
Abstract
The review by Reeg D [...]
Full article
Open AccessArticle
Brucellosis Seropositivity Using Three Serological Tests and Associated Risk Factors in Abattoir Workers in Gauteng Province, South Africa
by
, , , , , , , , and
Pathogens 2024, 13(1), 64; https://doi.org/10.3390/pathogens13010064 - 09 Jan 2024
Abstract
Abattoir workers are liable to zoonotic infections from animals and animal products, primarily to diseases with asymptomatic and chronic clinical manifestations in animals, such as brucellosis. No published reports exist on the seroprevalence of brucellosis in abattoir workers in South Africa. Therefore, this
[...] Read more.
Abattoir workers are liable to zoonotic infections from animals and animal products, primarily to diseases with asymptomatic and chronic clinical manifestations in animals, such as brucellosis. No published reports exist on the seroprevalence of brucellosis in abattoir workers in South Africa. Therefore, this cross-sectional study was conducted to estimate the occurrence and risk factors for Brucella exposure in abattoir workers in Gauteng Province. A total of 103 abattoir workers and managers from 6 abattoirs, where brucellosis-positive slaughtered cattle and sheep were previously detected, were interviewed and tested with serological assays using the Rose Bengal test (RBT), BrucellaCapt, and IgG-ELISA. A pre-tested questionnaire was administered to consenting respondents to obtain information on risk factors for brucellosis. Of the 103 respondents tested, the distribution of female and male workers was 16 (15.5%) and 87 (84.5%), respectively. The seroprevalence for exposure to brucellosis was 21/103 (20.4%, 95%CI: 13.1–29.5) using a combination of RBT, BrucellaCapt, or IgG-ELISA. For test-specific results, seroprevalences by RBT, BrucellaCapt, and IgG-ELISA were 13/103 (12.6%, 95%CI: 6.9–20.6), 9/103 (8.74%, 95%CI: 4.1–15.9), and 18/103 (17.5%, 95%CI: 10.7–26.2), respectively. Low-throughput abattoirs were identified as associated risks, as 29.3% of workers were seropositive compared with 12.7% of workers in high-throughput abattoirs, which highlights that direct contact at abattoirs poses higher risk to workers than indirect and direct contact outside abattoirs. This study confirms the occurrence of Brucella spp. antibodies among abattoir workers in South Africa, possibly due to occupational exposure to Brucella spp., and highlights the occupational hazard to workers. Furthermore, findings underscore that abattoir facilities can serve as points for active and passive surveillance for indicators of diseases of public health importance. We recommend periodic implementation of brucellosis testing of abattoir workers country-wide to establish baseline data for informing appropriate preventive practices and reducing the potential burden of infection rates among these high-risk workers.
Full article
(This article belongs to the Special Issue Bacterial Infections: Surveillance, Prevention and Control)
Open AccessCommunication
Genomic Features of an MDR Escherichia coli ST5506 Harboring an IncHI2/In229/blaCTX-M-2 Array Isolated from a Migratory Black Skimmer
by
, , , , , and
Pathogens 2024, 13(1), 63; https://doi.org/10.3390/pathogens13010063 - 09 Jan 2024
Abstract
Migratory birds have contributed to the dissemination of multidrug-resistant (MDR) bacteria across the continents. A CTX-M-2-producing Escherichia coli was isolated from a black skimmer (Rynchops niger) in Southeast Brazil. The whole genome was sequenced using the Illumina NextSeq platform and de
[...] Read more.
Migratory birds have contributed to the dissemination of multidrug-resistant (MDR) bacteria across the continents. A CTX-M-2-producing Escherichia coli was isolated from a black skimmer (Rynchops niger) in Southeast Brazil. The whole genome was sequenced using the Illumina NextSeq platform and de novo assembled by CLC. Bioinformatic analyses were carried out using tools from the Center for Genomic Epidemiology. The genome size was estimated at 4.9 Mb, with 4790 coding sequences. A wide resistome was detected, with genes encoding resistance to several clinically significant antimicrobials, heavy metals, and biocides. The blaCTX-M-2 gene was inserted in an In229 class 1 integron inside a ∆TnAs3 transposon located in an IncHI2/ST2 plasmid. The strain was assigned to ST5506, CH type fumC19/fimH32, serotype O8:K87, and phylogroup B1. Virulence genes associated with survival in acid conditions, increased serum survival, and adherence were also identified. These data highlight the role of migratory seabirds as reservoirs and carriers of antimicrobial resistance determinants and can help to elucidate the antimicrobial resistance dynamics under a One Health perspective.
Full article
(This article belongs to the Special Issue Detection and Epidemiology of Drug-Resistant Bacteria)
►▼
Show Figures
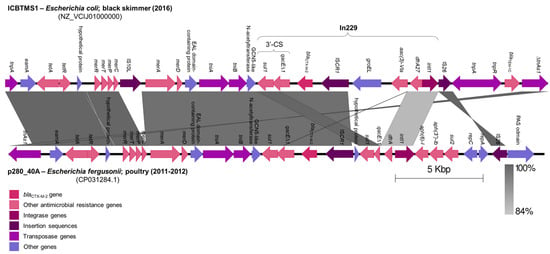
Figure 1
Open AccessFeature PaperArticle
Heat Inactivation of Nipah Virus for Downstream Single-Cell RNA Sequencing Does Not Interfere with Sample Quality
by
, , , , , , , , , , , and
Pathogens 2024, 13(1), 62; https://doi.org/10.3390/pathogens13010062 - 09 Jan 2024
Abstract
Single-cell RNA sequencing (scRNA-seq) technologies are instrumental to improving our understanding of virus–host interactions in cell culture infection studies and complex biological systems because they allow separating the transcriptional signatures of infected versus non-infected bystander cells. A drawback of using biosafety level (BSL)
[...] Read more.
Single-cell RNA sequencing (scRNA-seq) technologies are instrumental to improving our understanding of virus–host interactions in cell culture infection studies and complex biological systems because they allow separating the transcriptional signatures of infected versus non-infected bystander cells. A drawback of using biosafety level (BSL) 4 pathogens is that protocols are typically developed without consideration of virus inactivation during the procedure. To ensure complete inactivation of virus-containing samples for downstream analyses, an adaptation of the workflow is needed. Focusing on a commercially available microfluidic partitioning scRNA-seq platform to prepare samples for scRNA-seq, we tested various chemical and physical components of the platform for their ability to inactivate Nipah virus (NiV), a BSL-4 pathogen that belongs to the group of nonsegmented negative-sense RNA viruses. The only step of the standard protocol that led to NiV inactivation was a 5 min incubation at 85 °C. To comply with the more stringent biosafety requirements for BSL-4-derived samples, we included an additional heat step after cDNA synthesis. This step alone was sufficient to inactivate NiV-containing samples, adding to the necessary inactivation redundancy. Importantly, the additional heat step did not affect sample quality or downstream scRNA-seq results.
Full article
(This article belongs to the Special Issue Editorial Board Members’ Collection Series: Molecular Epidemiology, Diagnostics and Management of Viral Pathogens)
►▼
Show Figures
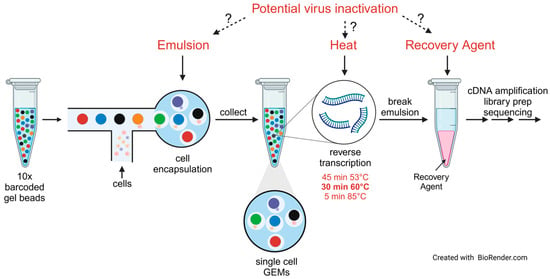
Figure 1
Open AccessArticle
NASCarD (Nanopore Adaptive Sampling with Carrier DNA): A Rapid, PCR-Free Method for SARS-CoV-2 Whole-Genome Sequencing in Clinical Samples
by
, , , , , , and
Pathogens 2024, 13(1), 61; https://doi.org/10.3390/pathogens13010061 - 09 Jan 2024
Abstract
Whole-genome sequencing (WGS) represents the main technology for SARS-CoV-2 lineage characterization in diagnostic laboratories worldwide. The rapid, near-full-length sequencing of the viral genome is commonly enabled by high-throughput sequencing of PCR amplicons derived from cDNA molecules. Here, we present a new approach called
[...] Read more.
Whole-genome sequencing (WGS) represents the main technology for SARS-CoV-2 lineage characterization in diagnostic laboratories worldwide. The rapid, near-full-length sequencing of the viral genome is commonly enabled by high-throughput sequencing of PCR amplicons derived from cDNA molecules. Here, we present a new approach called NASCarD (Nanopore Adaptive Sampling with Carrier DNA), which allows a low amount of nucleic acids to be sequenced while selectively enriching for sequences of interest, hence limiting the production of non-target sequences. Using COVID-19 positive samples available during the omicron wave, we demonstrate how the method may lead to >99% genome completeness of the SARS-CoV-2 genome sequences within 7 h of sequencing at a competitive cost. The new approach may have applications beyond SARS-CoV-2 sequencing for other DNA or RNA pathogens in clinical samples.
Full article
(This article belongs to the Special Issue New Insights into Molecular Epidemiology and Diagnostics of Human Pathogens)
►▼
Show Figures
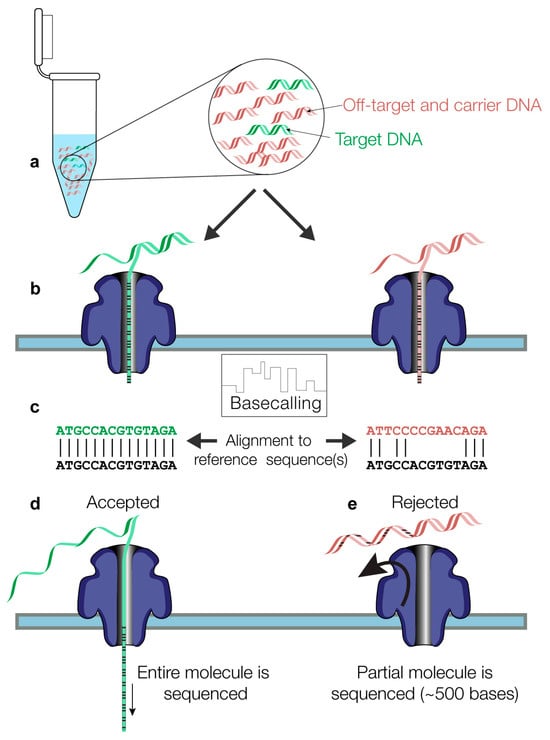
Figure 1
Open AccessReview
G-Quadruplexes in the Regulation of Viral Gene Expressions and Their Impacts on Controlling Infection
Pathogens 2024, 13(1), 60; https://doi.org/10.3390/pathogens13010060 - 08 Jan 2024
Abstract
G-quadruplexes (G4s) are noncanonical nucleic acid structures that play significant roles in regulating various biological processes, including replication, transcription, translation, and recombination. Recent studies have identified G4s in the genomes of several viruses, such as herpes viruses, hepatitis viruses, and human coronaviruses. These
[...] Read more.
G-quadruplexes (G4s) are noncanonical nucleic acid structures that play significant roles in regulating various biological processes, including replication, transcription, translation, and recombination. Recent studies have identified G4s in the genomes of several viruses, such as herpes viruses, hepatitis viruses, and human coronaviruses. These structures are implicated in regulating viral transcription, replication, and virion production, influencing viral infectivity and pathogenesis. G4-stabilizing ligands, like TMPyP4, PhenDC3, and BRACO19, show potential antiviral properties by targeting and stabilizing G4 structures, inhibiting essential viral life-cycle processes. This review delves into the existing literature on G4’s involvement in viral regulation, emphasizing specific G4-stabilizing ligands. While progress has been made in understanding how these ligands regulate viruses, further research is needed to elucidate the mechanisms through which G4s impact viral processes. More research is necessary to develop G4-stabilizing ligands as novel antiviral agents. The increasing body of literature underscores the importance of G4s in viral biology and the development of innovative therapeutic strategies against viral infections. Despite some ligands’ known regulatory effects on viruses, a deeper comprehension of the multifaceted impact of G4s on viral processes is essential. This review advocates for intensified research to unravel the intricate relationship between G4s and viral processes, paving the way for novel antiviral treatments.
Full article
(This article belongs to the Special Issue Host Immune Responses to RNA Viruses, Volume II)
►▼
Show Figures
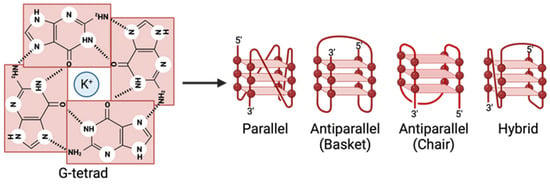
Figure 1
Open AccessArticle
Accurate Diagnosis of Schistosoma mansoni and S. haematobium from Filtered Urine Samples Collected in Tanzania, Africa
Pathogens 2024, 13(1), 59; https://doi.org/10.3390/pathogens13010059 - 08 Jan 2024
Abstract
Schistosomiasis is a bloodborne, and waterborne parasitic disease caused by the human Schistosoma species, namely Schistosoma mansoni and S. haematobium. The parasite requires an intermediate snail host, where they grow and develop, along with a human host (definitive). Schistosoma egg detection in
[...] Read more.
Schistosomiasis is a bloodborne, and waterborne parasitic disease caused by the human Schistosoma species, namely Schistosoma mansoni and S. haematobium. The parasite requires an intermediate snail host, where they grow and develop, along with a human host (definitive). Schistosoma egg detection in feces (S. mansoni) and urine (S. haematobium) are the WHO-recommended confirmatory diagnostic tests. The goal of our research was to determine the efficacy of detecting single or dual Schistosome species from filtered human urine samples collected in Tanzania by amplifying species-specific cell-free repeat DNA fragments via polymerase chain reaction (PCR) and gel electrophoresis. In total, 104 filtered human urine samples were evaluated and collected from individuals residing in the village of Kayenze, Tanzania. All samples were detected with 100% accuracy and no cross-amplification was present. For a single infection of S. mansoni, 22 (21%) of the samples were positive, while 15 (14%) of the samples were negative via PCR. Moreover, for a single infection of S. haematobium, 7 (7%) of the samples were positive, while 15 (14%) of the samples were negative. Dual infections were found in a higher percentage, with 60 (58%) of the samples being positive. Thus, we have justified that PCR is more sensitive and specific by amplifying species-specific cell-free repeat DNA fragments from the same urine sample than WHO-recommended methods of processing stool and urine.
Full article
(This article belongs to the Special Issue Epidemiology and Immunopathology of Intestinal Helminthiasis and Schistosomiasis: From Experimental Data to Field Work)
►▼
Show Figures
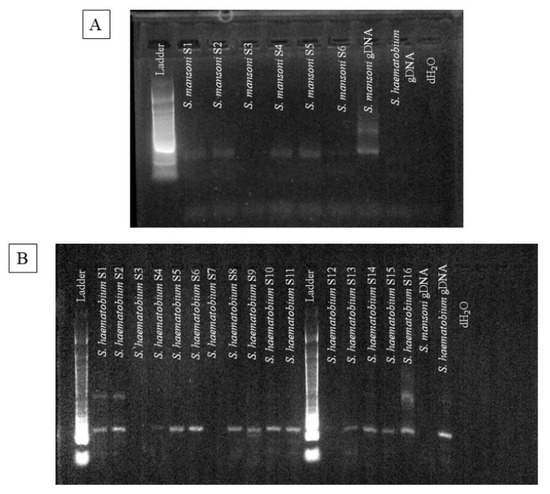
Figure 1

Journal Menu
► ▼ Journal Menu-
- Pathogens Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biomolecules, Cells, JMP, Pathogens, Viruses
Molecular and Cellular Mechanisms of Diseases: HIV
Topic Editors: Mara Biasin, Daria Lucia TrabattoniDeadline: 1 February 2024
Topic in
Brain Sciences, Cells, Diagnostics, IJMS, JCM, Pathogens, Pathophysiology
Applied Sciences and Technologies for Detection and Therapies of Pathologies in the Neuronal Environment
Topic Editors: Muh-Shi Lin, Hong Jiang, Yu-Yo SunDeadline: 29 February 2024
Topic in
Biology, Cancers, Current Oncology, Diseases, JCM, Pathogens
Pathogenetic, Diagnostic and Therapeutic Perspectives in Head and Neck Cancer
Topic Editors: Shun-Fa Yang, Ming-Hsien ChienDeadline: 20 June 2024
Topic in
Diseases, IJMS, Microbiology Research, Pathogens, Vaccines
Advances in Human Pathogen Control—a 21st Century Challenge 2.0
Topic Editors: Jorge H. Leitão, Nitin Amdare, Joana R FelicianoDeadline: 30 June 2024

Conferences
Special Issues
Special Issue in
Pathogens
Innovations and Developments in the Diagnosis of Sexually Transmitted Diseases
Guest Editors: Israel Nissan, Itzchak LevyDeadline: 20 January 2024
Special Issue in
Pathogens
Foot-and-Mouth Disease Virus: Pathogenesis and Persistence
Guest Editors: Michael Eschbaumer, Florian Pfaff, Christine LuttermannDeadline: 25 January 2024
Special Issue in
Pathogens
Exploring the Epidemiology, Pathogenicity, and Therapeutic Options of Staphylococcus spp.
Guest Editors: Francesca Paola Nocera, Patrizia NebbiaDeadline: 15 February 2024
Special Issue in
Pathogens
The Impact of Epstein Barr Virus (EBV) in Clinical Practice
Guest Editor: Fabio Salvatore MacalusoDeadline: 25 February 2024
Topical Collections
Topical Collection in
Pathogens
Mastitis in Dairy Ruminants
Collection Editors: George Fthenakis, Maria Filippa Addis
Topical Collection in
Pathogens
Feature Papers on Parasitic Pathogens
Collection Editor: Sébastien Besteiro
Topical Collection in
Pathogens
Emerging and Re-emerging Pathogens
Collection Editors: Sheng-Fan Wang, Wen-Hung Wang, Arunee Thitithanyanont
Topical Collection in
Pathogens
New Insights into Bacterial Pathogenesis
Collection Editor: Carmelo Biondo









.jpg)







